Diet-Induced Alterations in Gut Microbiota and Antimicrobial Peptides Profiles of Honey Bees (Apis mellifera)
Abstract
Honey bees (Apis mellifera) are essential not only for honey production but also as key pollinators in agriculture. Worker bees perform age-related tasks, remaining inside the hive for approximately the first 20 days after emergence and foraging outside thereafter, which is critical for colony maintenance. However, complex stressors weaken the bee’s immune system and disrupt its gut microbiome. Given that the gut microbiome plays an essential role in immune defense, this disruption renders honey bees susceptible to pathogens, contributing to colony collapse disorder. In this study, we investigated the effects of larval and adult diet on gut microbial communities and antimicrobial peptides (AMPs) expression in worker bees over 20 days. Each group was initially assigned to one of two dietary treatments: a 50% sucrose solution or honey. The adult individuals that developed under these conditions were subsequently maintained on their respective initial diets, thereby generating four experimental groups. At the end of the feeding period, gut microbiota composition and the expression of AMPs were quantified using real-time quantitative PCR. Additionally, cultivable gut bacteria were isolated and identified for comparative analysis. Our results show that feeding honey reduced harmful bacteria and increased beneficial bacteria associated with AMPs expression. These results suggest that diet during development can improve gut health and immunity in honeybees.
Keywords:
Apis mellifera, Colony Collapse Disorder, Gut microbiota, Dietary influence, Antimicrobial peptides expressionINTRODUCTION
Animal pollinators, including both wild and domesticated bees, are increasingly recognized as the primary agents of pollination, providing essential services to wild flora and economically important crops within natural and agricultural ecosystems (Gallai et al., 2009; Papa et al., 2022). In particular, managed colonies of honey bees (Apis mellifera) play a disproportionately significant role; in the United States alone, they contributed an estimated $11.7 billion in pollination services as of 2009 (Powell et al., 2021; Moharrami et al., 2022). Bees of the Apidae family are recognized as the most essential pollinators worldwide, with the domesticated European honeybee, A. mellifera, responsible for delivering nearly 80% of global agricultural pollination services (Carreck and Williams, 1998; Formato et al., 2024). This critical role not only sustains crop production but also underpins biodiversity and the overall health of ecosystems.
In recent years, certain regions have seen an increase in honey bee colony losses, with Colony Collapse Disorder (CCD)—first reported in the United States in 2006—identified as a primary cause (Neumann and Carreck, 2010). CCD is characterized by the sudden disappearance of adult worker bees, the absence of significant numbers of dead bees, and a delayed onset of pest infestations or robbing. In the final stages, only the queen and a small number of workers remain, while capped brood and stored food are left intact (Cox-Foster et al., 2007; VanEngelsdorp et al., 2009; Williams et al., 2010). Although the exact cause of CCD has not yet been determined, it is widely accepted that the phenomenon results from the combined effects of various environmental stressors, including pesticide exposure, Varroa mite infestation, viral pathogens, climatic changes, and the gradual loss of natural habitats (Neov et al., 2019). Such adverse factors compromise the immune competence of honey bees and perturb the gut microbiota, which serves as a pivotal determinant of host health and defense against pathogenic threats (Rosenkranz et al., 2010; Flores et al., 2021). The functional benefits conferred by the native gut microbiome of honey bees extend well beyond mere protection from pathogenic organisms. Growing empirical evidence indicates that the gut microbiota of social bees mediates colonization resistance, thereby limiting the establishment and proliferation of potentially harmful microorganisms (Maes et al., 2016; Raymann et al., 2017). In addition to this protective role, recent studies have shown that the native gut microbiota actively stimulates the host immune system by inducing the expression of antimicrobial peptides (AMPs) such as Apidaecin and Hymenoptaecin (Kwong et al., 2017). This dual function-direct inhibition of pathogen colonization and immune modulation-highlights the gut microbiome as a critical determinant of colony health.
Diverse carbohydrate sources, such as sucrose, invert sugar, and components of honey, directly impact the composition of core gut bacteria in adult bees. Notably, natural food sources like honey have been shown to increase the proportion of beneficial bacteria, thereby enhancing gut microbial balance. This diet-induced modulation of the gut microbiome is closely linked to the host’s resistance to infectious diseases (Harris et al., 2019; Taylor et al., 2019). For example, nutritionally poor diets can disrupt the honeybee’s gut microbial equilibrium and down-regulate the expression of immune-related genes, consequently increasing susceptibility to pathogens like Nosema ceranae (Castelli et al., 2020). These findings collectively suggest that diet is a key determinant of honeybee gut health, with downstream effects on immune function and disease defense mechanisms.
This study evaluated the association between changes in gut microbiota composition and expression of AMPs in worker honey bees subjected to dietary variations during different developmental stages. Given the current deficiency of research on the effects of diet across the honey bee life cycle on gut microbial community structure, the findings are expected to contribute to research aimed at improving honey bee immunity.
MATERIALS AND METHODS
1. Experimental procedures
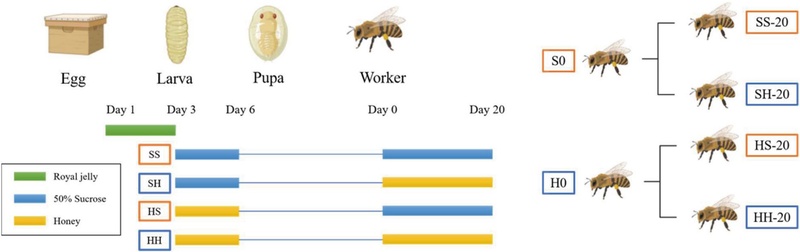
To investigate the effects of diet on gut microbial composition, dietary regimens were controlled separately during the larval, pupa, and first 20 days of the adult stage. The experiment was conducted according to the method described by Lanh et al. (2022). Apis mellifera larvae were sourced from the experimental apiary at the College of Ecological and Environmental Science, Kyungpook National University, Sangju-si, Republic of Korea. During the first three days post-hatching, larvae were exclusively provisioned with royal jelly. From the fourth day of the larval stage until adult eclosion, individuals were assigned to the two distinct dietary treatments: a 50% (w/v) sucrose solution (group S0) or natural honey (group H0). Upon emergence, adult workers were subdivided into four experimental groups, each comprising 20 individuals, and assigned feeding treatments consistent with their larval-stage diet, yielding the following groups: SS (sucrose-sucrose), SH (sucrose-honey), HS (honey-sucrose), and HH (honey-honey). A schematic overview of the experimental design is depicted in Fig. 1. Both larval and adult bees were incubated at 32℃ in a temperature-controlled incubator, and fresh feed was provided daily in the morning.
2. Gut microbial composition analysis
Three individual bees from each group were washed in 70% ethanol, and the heads and wings were removed before being stored at -80℃. Following homogenization using a Bullet Blender (Bertin Technologies, Montigny-le-Bretonneux, France), genomic DNA were simultaneously extracted from frozen samples with the Clear-STM DNA/RNA extraction kit (INVIRUSTECH, Gwangju, Korea). The purity and concentration of the extracted DNA were subsequently assessed using a SpectraMax® QuickDropTM UV-Vis Spectrophotometer (Molecular Devices, Sunnyvale, CA, USA).
To analyze the gut microbiota composition of honey bees in relation to their dietary sources, bacterial species identified as beneficial, pathogenic, or opportunistic were systematically categorized using data retrieved from the National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov). Primers targeting the 16S rRNA genes of each bacterial group were designed as shown in Table 1 (Mattila et al., 2012). PCR amplification was performed using 100 ng of genomic DNA as template, with 10 pmole of each forward and reverse primer per reaction. The amplification conditions were as follows: an initial denaturation at 95℃ for 1 minute, followed by 40 cycles consisting of denaturation at 95℃ for 15 seconds, annealing at 55℃ for 15 seconds, and extension at 72℃ for 30 seconds (Moškrič et al., 2023).
3. cDNA synthesis and quantitative PCR analysis
For each experimental group, three individual bees were used as biological replicates. Total RNA was extracted using the yesRTM total RNA extraction kit with a gDNA Eliminator column (Prefilter PF02) (GenesGen, Busan, Korea). Complementary DNA synthesis was performed with ReverTra Ace reverse transcriptase following the manufacturer’s recommended protocol (Toyobo, Osaka, Japan). Specific primers targeting the AMPs (Abaecin, Apidaecin, Defensin1, Defensin2, and Hymenoptaecin) were designed for expression analysis in honeybees with RPS5 gene (Kim et al., 2022) transcript as a reference (Table 2). Real-time quantitative PCR reaction contained 200 ng of template DNA and 10 pmole of each forward and reverse primer. After an initial pre-denaturation step at 95℃ for 1 minute, amplification was carried out for 40 cycles of 95℃ for 15 seconds, 55℃ for 15 seconds, and 72℃ for 30 seconds to determine gene expression levels. The experiments were performed in triplicate under the same conditions.
4. Gut microbiota isolation and identification
For each of the S0 and H0 groups, ten bees were prepared by washing them on ice at 4℃ with 20mL of 70% ethanol, followed by a rinse with phosphate-buffered saline. After removal of the head, wings, and esophagus, the digestive and excretory organs were dissected, homogenized, and centrifuged at 4,000 rpm for 5 minutes to collect the supernatant. The obtained supernatant was diluted, and 100μL aliquots were plated onto Luria-Bertani (LB; Difco, Sparks, MD, United States) agar and de Man, Rogosa and Sharpe (MRS; Difco, Detroit, MI, USA) agar media. To detect various lactic acid bacteria, MRS agar was supplemented with 0.02% bromocresol purple (BPB), and colony color changes due to lactic acid production were observed. The plates were incubated at 37℃ for 24 hours, with the incubation period extended as needed until visible colonies appeared. Colonies from each medium were then isolated and sub-cultured under the same conditions to obtain pure single colonies (Audisio et al., 2011).
Genomic DNA for bacterial identification was extracted from samples employing the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) following the manufacturer’s instructions. For PCR amplification, 100 ng of genomic DNA from each bacterial isolate and 10 pmole of the universal bacterial primers 27F (forward; 5ʹ-AGAGTTTGATCMTGGCTCAG-3ʹ) and 1492R (reverse; 5ʹ-GGTTACCTTGTTACGACTT-3ʹ) were combined with the AccuPower® PCR PreMix & Master Mix (Bioneer, Daejeon, Korea). The cycling conditions included an initial denaturation at 95℃ for 2 minutes, followed by 22 cycles of 94℃ for 30 seconds, 55℃ for 30 seconds, and 72℃ for 1 minutes, with a final extension step at 72℃ for 5 minutes (Koch and Schmid-Hempel, 2013). Amplification was confirmed by electrophoresis on a 1.5% agarose gel. The PCR products were purified and sequenced by Solgent (http://www.solgent.com). Microbial identification was performed using the BLAST search program within the NCBI database (https://www.ncbi.nlm.nih.gov).
5. Statistical analysis
One-way analysis of variance was performed using SPSS software (version 24, SPSS, Inc., Chicago, IL, USA). The mean values were compared using Scheffé’s method, and a p value<0.05 was considered significant.
RESULTS
1. Changes in gut microbiota induced by different feed sources
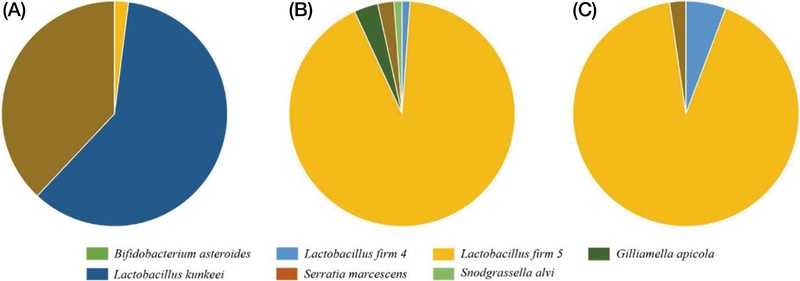
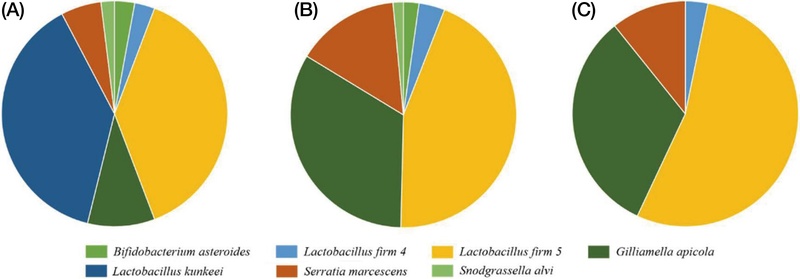
Analysis of gut microbial composition in the S0 group, which was provided with the 50% sucrose solution during the larval stage, demonstrated that L. kunkeei predominated, while the L. firm 5 cluster was present at minimal levels. Moreover, S. marcescens, recognized as an opportunistic pathogen in honey bees, was identified. In contrast, the SS group showed a increase in the relative abundance of L. firm 5, alongside the emergence of G. apicola and S. alvi, which was associated with a decreased proportion of S. marcescens (Fig. 2).
During the same period, the H0 group, which was fed natural honey, exhibited an approximately balanced ratio of L. kunkeei to L. firm 5 (about 1 : 1), with the additional presence of L. firm 4. The relative abundance of S. marcescens was markedly lower in the H0 group compared to the S0 group. Furthermore, strains known to be part of the honey bee gut microbiota, such as B. asteroides, were also detected. In the HS and HH groups (Fig. 2), L. firm 5 was identified as the predominant lactic acid bacterium among the previously balanced L. kunkeei and L. firm 5 populations observed in the H0 group. However, an increasing trend in the proportion of potential pathogenic bacteria was observed, with specifically higher levels detected in groups administered 50% sucrose solution (Fig. 3).
2. Dietary effects on AMP related genes
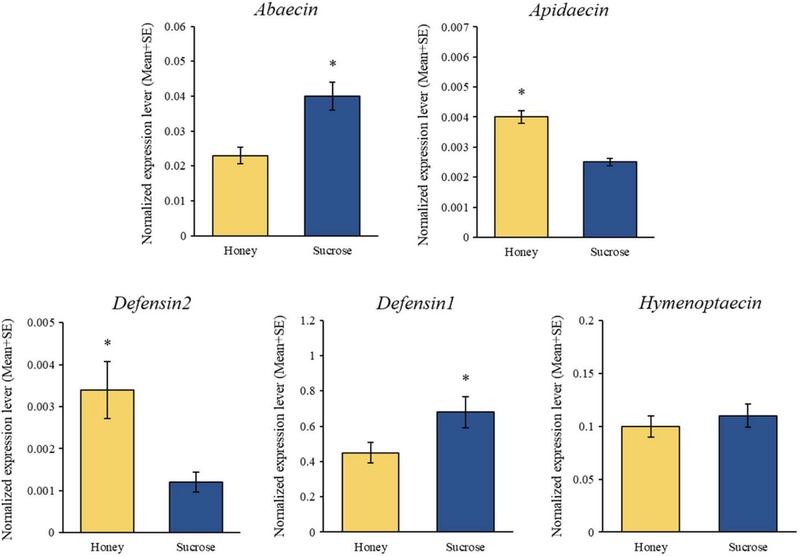
The expression levels of five honey bee AMPs (Abaecin, Apidaecin, Defensin1, Defensin2, and Hymenoptaecin) were compared between the S0 and H0 groups. Apidaecin and Defensin2 exhibited higher expression levels in the H0 group, whereas Abaecin and Defensin1 were more highly expressed in the S0 group. Notably, Defensin2 expression in the S0 group was more than twice that observed in the H0 group. No significant difference in Hymenoptaecin expression was detected between the two groups (Fig. 4).
3. Comparative analysis of cultivable gut microbiota
Given the observed reduction of pathogenic bacteria in both the SS and SH groups, we isolated and identified their gut microbiota to compare the compositions of beneficial and pathogenic bacteria. The results showed that, at the same dilution factors, the colony-forming units of pathogenic bacteria were lower in the SH group. Furthermore, enhanced proliferation of lactic acid bacteria was observed in the SH group on MRS medium supplemented with 0.02% BPB.
In the SS group, the isolates were predominantly Gram-negative bacteria, including Escherichia coli and Enterobacter aerogenes. On MRS agar, non-lactic acid bacteria from the SS group were identified as Gibbsiella sp., while a Lactobacillus sp. was detected at low frequency. In contrast, isolates from the SH group were identified as Bacillus sp., which are known to be beneficial to honeybees. Most colonies on MRS agar were Lactobacillus kunkeei, and the BPB-negative colonies were identified as Bacillus sp. (Table 3).
DISCUSSION
In this study, we investigated the influence of dietary sources on the formation and composition of the gut microbial community and characterized its taxonomic features. The importance of microbiomes lies not only in their interaction with the host but also in their composition, metabolites, and gene products, which are modulated by the host’s environment and lifestyle, particularly diet. A diverse diet increases the abundance of symbionts, thereby benefiting the host through its effects on biological processes, behavioral patterns, developmental pathways, and immune responses (Yun et al., 2014; Krams et al., 2017). The abdominal gut of the honeybee is anatomically divided into three distinct regions-the foregut, midgut, and hindgut-each harboring characteristic and relatively stable bacterial communities (Raymann and Moran, 2018). The foregut, comprising the esophagus and crop, exhibits low microbial density and is highly responsive to dietary composition. In the midgut, bacteria predominantly attach to the epithelial lining, with honeybee-specific taxa such as G. apicola and S. alvi being prevalent. At the junction between the midgut and hindgut lie the filamentous Malpighian tubules, where F. perrara has been reported. The hindgut harbors the highest density of gut bacteria, predominantly lactic acid bacteria from the Firm 4 and Firm 5 clades (Lactobacillus spp.) as well as Bifidobacterium spp. (Engel et al., 2012; Martinson et al., 2012; Kwong and Moran, 2016). Within the hindgut, the rectum is dominated by L. Firm 5, L. Firm 4, and B. asteroides, whereas in the ileum, S. alvi, G. apicola, and L. Firm 5 are predominant. Consistent with these reports, the present study identified Snodgrassella, Gilliamella, and diverse species of Lactobacillus and Bifidobacterium as core members of the honeybee gut microbiota, supporting the structural universality and functional stability of these communities. However, it is important to note that our qPCR-based quantification of bacterial taxa relied on targeting the 16S rRNA gene. The copy number of this gene can vary significantly among different bacterial species and strains (Vĕtrovský and Baldrian, 2013). Therefore, the relative abundance data presented may not perfectly reflect the true cell-based community composition. However, as the primary conclusions of this study are based on the relative comparison of bacterial loads between treatment and control groups, this inherent limitation is unlikely to alter the interpretation of our key findings. Future studies could employ copy number correction using databases like rrnDB to achieve more precise quantification (Stoddard et al., 2015).
Beneficial bacteria within the honeybee gut produce metabolic byproducts that reduce intestinal pH, thereby inhibiting the proliferation of pathogenic bacteria (Daisley et al., 2020a). They also occupy ecological niches in the gut, competitively excluding colonization by pathogenic microbes, and contribute to the detoxification of dietary toxins and environmental stressors, thus supporting overall host health (Daisley et al., 2020b). Furthermore, these beneficial microbes induce the expression of AMPs thereby enhancing systemic immune function in honeybees (Chege et al., 2023; Motta and Moran, 2024). The honeybee gut symbiont S. alvi promotes the expression of apidaecin and hymenoptaecin, thereby enhancing defense against pathogenic microorganisms. Lactobacillus spp. stimulates the expression of defensin1 and defensin2, contributing to the stabilization of the gut environment and strengthening host immunity (Lang et al., 2022), while G. apicola also modulates AMPs expression through interactions with the host innate immune system. The expression of these AMPs is regulated via the Toll and Imd signaling pathways, which are closely associated with gut microbial community composition, and collectively play a pivotal role in maintaining honeybee health and resistance to pathogens (Engel et al., 2016; Emery et al., 2017). The lactic acid bacterium Lactobacillus kunkeei, which is known for its ability to stimulate AMPs expression and produce bacteriocins (Zendo et al., 2020), was also detected in this study. However, its abundance was observed to decline markedly in adult bees after 20 days of consistent feeding on the same diet. This decline suggests a dynamic microbial response to long-term dietary conditions that warrants further investigation. Furthermore, a key limitation of this study is that AMPs expression was assessed only in the S0 and H0 groups. These findings suggest the need for two separate follow-up experiments: (1) an independent investigation to determine the causes of the reduction of L. kunkeei and other lactic acid bacteria and their impact on gut microbial community dynamics, and (2) a more comprehensive analysis of AMPs expression across multiple conditions and time points.
Research on the interaction between gut microbiota and honeybee immunity provides critical insights into honeybee health and resistance to pathogens, contributing significantly to the preservation of bee ecosystems and the sustainability of apiculture. In this study, we confirmed that dietary type influences the composition of the gut microbial community, which in turn stimulates the expression of AMP-related factors. These findings suggest that the results of this work could serve as important foundational data for the discovery of potential probiotic strains for honeybees. Nevertheless, further investigations are required to clarify the causal relationship between changes in lactic acid bacterial populations and immune gene expression, which could ultimately enable the development of tailored microbiome management strategies to enhance honeybee immunity and overall health. In addition, the methodological approach of this study, which relied exclusively on aerobic culturing, presents an inherent limitation. The isolation protocol was designed specifically to recover aerobic and facultatively anaerobic bacteria. Consequently, the collection of isolates reported herein does not represent the complete microbial diversity of the honeybee gut, a predominantly anaerobic environment. Therefore, the isolates obtained in this study should be interpreted as a specific, culturable subset of the total gut community, rather than a comprehensive census. To overcome this methodological constraint and achieve a holistic understanding of the community’s compositional dynamics in response to diet, the integration of culture-independent techniques is required. Future investigations should incorporate a metagenomic approach to characterize the full spectrum of microbial diversity. Such analyses would provide an essential framework for contextualizing the ecological and functional roles of the specific isolates recovered in the present study, thereby enabling a more complete elucidation of the relationship between diet, the gut microbiome, and honeybee health.
CONCLUSION
Based on the results of this study, our findings demonstrate that larval diet composition exerts a critical influence on gut microbial structure and immune-related gene expression in honeybees. A honey-based diet promoted the proliferation of beneficial taxa, including Bacillus spp. and Lactobacillus kunkeei, which were strongly correlated with enhanced immune responses. Specifically, when adult bees that were fed a 50% sucrose diet during their larval stage were subsequently given honey, the proportion of opportunistic pathogens decreased, while beneficial bacteria increased, successfully restoring the disturbed gut microbial ecosystem. These results indicate that the detrimental effects of artificial sugar diets may be reversible and underscore the essential role of natural honey in re-establishing and sustaining honeybee health. Further research is needed to clarify the underlying mechanisms connecting dietary interventions to immune modulation in adult honeybees.
Acknowledgments
This study was supported by a research fund (RS-2025-02303308) from the Rural Development Administration, Republic of Korea.
References
-
Audisio, M. C., D. Torres and R. C. Fadda. 2011. Properties of different lactic acid bacteria isolated from Apis mellifera L. bee-gut. Microbiol. Res. 166(1): 1-13.
[https://doi.org/10.1016/j.micres.2010.01.003]

-
Carreck, N. and I. Williams. 1998. The economic value of bees in the UK. Bee World 79(3): 115-123.
[https://doi.org/10.1080/0005772X.1998.11099393]

-
Castelli, L., V. Tosi, F. Simonetti, G. Bordi and G. Piana. 2020. Impact of nutritional stress on honeybee gut microbiota, immunity, and Nosema ceranae infection. Microb. Ecol. 80(4): 908-919.
[https://doi.org/10.1007/s00248-020-01538-1]

-
Chege, M., J. Kinyua and J. C. Paredes. 2023. Lactobacillus kunkeei impacts the health of honey bees, Apis mellifera scutellata, and protects the bees against the opportunistic pathogen Serratia marcescens. Int. J. Trop. Insect Sci. 43(6): 1947-1955.
[https://doi.org/10.1007/s42690-023-01103-6]

-
Cox-Foster, D. L., S. Conlan, E. C. Holmes, G. E. Palacios, J. D. Evans and N. A. Moran. 2007. A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318(5848): 283-287.
[https://doi.org/10.1126/science.1146498]

-
Daisley, B. A., A. P. Pitek, J. A. Chmiel, S. Gibbons, A. M. Chernyshova, K. F. Al, K. M. Faragalla, J. P. Burton, G. J. Thompson and G. Reid. 2020a. Lactobacillus spp. attenuate antibiotic-induced immune and microbiota dysregulation in honey bees. Commun. Biol. 3(1): 534.
[https://doi.org/10.1038/s42003-020-01259-8]

-
Daisley, B. A., J. A. Chmiel, A. P. Pitek, G. J. Thompson and G. Reid. 2020b. Missing microbes in bees: how systematic depletion of key symbionts erodes immunity. Trends Microbiol. 28(12): 1010-1021.
[https://doi.org/10.1016/j.tim.2020.06.006]

-
Emery, O., K. Schmidt and P. Engel. 2017. Immune system stimulation by the gut symbiont Frischella perrara in the honey bee (Apis mellifera). Mol. Ecol. 26(9): 2576-2590.
[https://doi.org/10.1111/mec.14058]

-
Engel, P., V. G. Martinson and N. A. Moran. 2012. Functional diversity within the simple gut microbiota of the honey bee. Proc. Natl. Acad. Sci. U.S.A. 109(27): 11002-11007.
[https://doi.org/10.1073/pnas.1202970109]

-
Engel, P., W. K. Kwong and N. A. Moran. 2016. The bee microbiome: impact on bee health and model for evolution and ecology of host-microbe interactions. mBio 7(2): e02164-15.
[https://doi.org/10.1128/mBio.02164-15]

-
Flores, J. M., E. García-Vázquez and M. Martín-Hernández. 2021. Impact of Varroa destructor and associated pathologies on the colony collapse disorder affecting honey bees. Res. Vet. Sci. 135: 85-95.
[https://doi.org/10.1016/j.rvsc.2021.01.001]

-
Formato, G., G. Cilia, M. Zaccardelli and A. D’Antonio. 2024. Apis mellifera welfare: definition and future directions. Front. Anim. Sci. 5: 1486587.
[https://doi.org/10.3389/fanim.2024.1486587]

-
Gallai, N., J. M. Salles, J. Settele and B. E. Vaissière. 2009. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 68(3): 810-821.
[https://doi.org/10.1016/j.ecolecon.2008.06.014]

-
Harris, E. V., J. C. de Roode and N. M. Gerardo. 2019. Diet-microbiome-disease: investigating diet’s influence on infectious disease resistance through alteration of the gut microbiome. PLoS Pathog. 15(10): e1007891.
[https://doi.org/10.1371/journal.ppat.1007891]

-
Kim, Y., H. Kim, H. Cha, S. H. Lee and Y. H. Kim. 2022. Validation of quantitative real-time PCR reference genes and spatial expression profiles of detoxication-related genes under pesticide induction in honey bee, Apis mellifera. PLoS One 17(11): e0277455.
[https://doi.org/10.1371/journal.pone.0277455]

-
Koch, H. and P. Schmid-Hempel. 2013. Diversity and evolutionary patterns of bacterial gut associates of corbiculate bees. Mol. Ecol. 22(7): 2028-2044.
[https://doi.org/10.1111/mec.12209]

-
Krams, I. A., R. Krams and T. Krama. 2017. Microbiome symbionts and diet diversity incur costs on the immune system of insect larvae. J. Exp. Biol. 220(22): 4204-4212.
[https://doi.org/10.1242/jeb.169227]

-
Kwong, W. K. and N. A. Moran. 2016. Gut microbial communities of social bees. Nat. Rev. Microbiol. 14(6): 374-384.
[https://doi.org/10.1038/nrmicro.2016.43]

-
Kwong, W. K., A. L. Mancenido and N. A. Moran. 2017. Immune system stimulation by the native gut microbiota of honey bees. R. Soc. Open Sci. 4(2): 170003.
[https://doi.org/10.1098/rsos.170003]

-
Lang, H., Y. Wang, F. Zhu and X. Shen. 2022. Specific strains of honeybee gut Lactobacillus stimulate host immune system to protect against pathogenic Hafnia alvei. Microbiol. Spectr. 10(1): e01896-21.
[https://doi.org/10.1128/spectrum.01896-21]

-
Lanh, P. T., P. L. Nguyen and Q. D. Truong. 2022. The gut microbiota at different developmental stages of Apis cerana reveals potential probiotic bacteria for improving honeybee health. Microorganisms 10(10): 1938.
[https://doi.org/10.3390/microorganisms10101938]

-
Maes, P. W., L. Solter and K. Crailsheim. 2016. Diet-related gut bacterial dysbiosis correlates with impaired development, increased mortality and Nosema disease in the honeybee (Apis mellifera). Mol. Ecol. 25(21): 5439-5450.
[https://doi.org/10.1111/mec.13862]

-
Martinson, V. G., J. Moy and N. A. Moran. 2012. Establishment of characteristic gut bacteria during development of the honeybee worker. Appl. Environ. Microbiol. 78(8): 2830-2840.
[https://doi.org/10.1128/AEM.07810-11]

-
Mattila, H. R., D. Goulson and L. A. O’Toole. 2012. Characterization of the active microbiotas associated with honey bees reveals healthier and broader communities when colonies are genetically diverse. PLoS One 7(3): e32962.
[https://doi.org/10.1371/journal.pone.0032962]

- Moharrami, M., S. A. G. Zadeh and A. A. Aivazi. 2022. Role of honey bee gut microbiota in the control of American foulbrood and European foulbrood diseases. Arch. Razi Inst. 77(4): 1331.
-
Moškrič, A., A. Škorjanc and N. Ocvirk. 2023. Cutting corners: The impact of storage and DNA extraction on quality and quantity of DNA in honeybee (Apis mellifera) spermatheca. Front. Physiol. 14: 1139269.
[https://doi.org/10.3389/fphys.2023.1139269]

-
Motta, E. V. S. and N. A. Moran. 2024. The honeybee microbiota and its impact on health and disease. Nat. Rev. Microbiol. 22(3): 122-137.
[https://doi.org/10.1038/s41579-023-00990-3]

-
Neov, B., A. Al-Ghamdi, N. A. Al-Mekhlafi, H. R. El-Seedi, H. R and S. G. Musharraf. 2019. Biotic and abiotic factors associated with colonies mortalities of managed honey bee (Apis mellifera). Diversity 11(12): 237.
[https://doi.org/10.3390/d11120237]

-
Neumann, P. and N. L. Carreck. 2010. Honey bee colony losses. J. Apic. Res. 49(1): 1-6.
[https://doi.org/10.3896/IBRA.1.49.1.01]

-
Papa, G., G. Benelli and A. Cini. 2022. The honey bee Apis mellifera: an insect at the interface between human and ecosystem health. Biology 11(2): 233.
[https://doi.org/10.3390/biology11020233]

-
Powell, J. E., S. A. Ghabrial and M. R. Liles. 2021. Field-realistic tylosin exposure impacts honey bee microbiota and pathogen susceptibility, which is ameliorated by native gut probiotics. Microbiol. Spectr. 9(1): e0010321.
[https://doi.org/10.1128/Spectrum.00103-21]

-
Raymann, K. and N. A. Moran. 2018. The role of the gut microbiome in health and disease of adult honey bee workers. Curr. Opin. Insect Sci. 26: 97-104.
[https://doi.org/10.1016/j.cois.2018.02.012]

-
Raymann, K., Z. Shaffer and N. A. Moran. 2017. Antibiotic exposure perturbs the gut microbiota and elevates mortality in honeybees. PLoS Biol. 15(3): e2001861.
[https://doi.org/10.1371/journal.pbio.2001861]

-
Rosenkranz, P., P. Aumeier and B. Ziegelmann. 2010. Biology and control of Varroa destructor. J. Invertebr. Pathol. 103: S96-S119.
[https://doi.org/10.1016/j.jip.2009.07.016]

-
Stoddard, S. F., B. J. Smith, R. Hein, B. R. Roller and T. M. Schmidt. 2015. rrn DB: improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 43(D1): D593-D598.
[https://doi.org/10.1093/nar/gku1201]

-
Taylor, M. A., K. H. Lih, O. G. Baines and H. L. Mitchell. 2019. The effect of carbohydrate sources: Sucrose, invert sugar and components of mānuka honey, on core bacteria in the digestive tract of adult honey bees(Apis mellifera). PLoS One 14(12): e0225845.
[https://doi.org/10.1371/journal.pone.0225845]

-
VanEngelsdorp, D., J. D. Evans, C. Saegerman, C. Mullin, E. Haubruge and B. K. Nguyen. 2009. Colony collapse disorder: a descriptive study. PLoS One 4(8): e6481.
[https://doi.org/10.1371/journal.pone.0006481]

-
Vĕtrovský, T. and P. Baldrian. 2013. The variability of the 16S rRNA gene in bacterial genomes and its consequences for bacterial community analyses. PLoS One 8(2): e57923.
[https://doi.org/10.1371/journal.pone.0057923]

-
Williams, G. R., D. R. Tarpy, D. VanEngelsdorp and P. Neumann. 2010. Colony collapse disorder in context. Bioessays 32(10): 845.
[https://doi.org/10.1002/bies.201000075]

-
Yun, J., K. Kim, S. Park and Y. Kim. 2014. Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl. Environ. Microbiol. 80(17): 5254-5264.
[https://doi.org/10.1128/AEM.01226-14]

-
Zendo, T., M. Saimura, S. Shioya and M. Sugiyama. 2020. Kunkecin A, a new nisin variant bacteriocin produced by the fructophilic lactic acid bacterium, Apilactobacillus kunkeei FF30-6 isolated from honey bees. Front. Microbiol. 11: 571903.
[https://doi.org/10.3389/fmicb.2020.571903]