
Screening the Candidate Postmortem Marker Genes for Estimating Postmortem Intervals in Nurse Bees (Apis mellifera)
Abstract
The honey bee (Apis mellifera) plays an essential role in global agriculture; however, colony losses have been frequently reported. Because several mRNAs remain active for a limited period after death, quantitative real-time PCR(qRT-PCR) analysis of mRNA degradation may be useful for estimating postmortem intervals(PMIs) in dead bees. In this study, the expression dynamics of thirteen candidate postmortem marker genes were analyzed in nurse bee carcasses from 1 to 7 days postmortem. Among them, RPS5 and RPS18 showed significant decreases in expression within the early postmortem phase (1-3 days). Based on cubic regression models, RPS5 (R2= 0.997) and RPS18 (R2= 0.982) demonstrated strong correlations with PMI. Furthermore, expression ratios using stably expressed genes as denominators revealed that RPS5/SDH and RPS18/SDH achieved the highest coefficients of determination (R2= 0.996), distinguishing carcasses between 1-2 and 3-7 days postmortem. These findings suggest that RPS5, RPS18, and their ratios with SDH can serve as molecular markers for PMI estimation in nurse bees. This molecular approach may support forensic analyses of honey bee mortality under both laboratory and field conditions.
Keywords:
Apis mellifera, Postmortem interval, mRNA degradation, RPS5, RPS18, SDHINTRODUCTION
The western honey bee (Apis mellifera) is one of the most important pollinators contributing to agricultural productivity and global ecosystem stability. The Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services (IPBES) reported that approximately 35% of global crop production depends on pollination services, with an estimated economic value of 235-577 billion USD (Jung, 2008; IPBES, 2016). In Korea, honey bees are responsible for pollinating nearly 50% of fruit and vegetable crops, providing pollination services valued at approximately 4 billion USD (Jung, 2008). Similarly, Reilly et al.(2020) estimated that honey bee-mediated pollination contributes about 6.4 billion USD annually to U.S. crop yields, with almonds alone accounting for 4.2 billion USD.
However, these contributions have declined in parallel with the global reduction in honey bee populations (Sumner and Boriss, 2006; Meixner, 2010). Multiple factors-including exposure to pesticides and miticides, parasitic infestation, pathogens, and abnormal climatic conditions-have been identified as major stressors that threaten colony health and contribute to colony collapse (Meixner, 2010; Kim, 2022; Lee et al., 2022). Consequently, numerous studies have focused on the physiological effects of these stressors on surviving bees to identify potential lethal mechanisms (Zaobidna et al., 2017; Dolezal and Toth, 2018; Butolo et al., 2021; You et al., 2025).
Nevertheless, beekeepers often recognize colony losses only after large-scale mortality events occur, at which point most individuals are already dead. Thus, analyzing dead bees rather than survivors may provide more practical insight into the underlying causes of colony collapse. Following death, cellular metabolism ceases and autolytic degradation begins (Brooks and Sutton, 2017; Wenzlow et al., 2023). However, several studies have demonstrated that certain messenger RNAs (mRNAs) remain detectable for a limited period after death (Kimura et al., 2011; Pozhitkov et al., 2017; Shafeeq et al., 2020) reported that stress- and development-related genes continued to be expressed for 48-96 hours in postmortem mouse and zebrafish tissues, whereas Kimura et al. (2011) observed that circadian genes in mice maintained rhythmic expression patterns for up to 48 hours after death. In insects, Shafeeq et al. (2020) found that hsp70 expression increased for five days postmortem in Plodia interpunctella, while ultraspiracle transcripts remained stable.
Because the persistence of mRNA varies depending on the postmortem interval (PMI), these molecular changes have been proposed as biomarkers for estimating PMI (Scrivano et al., 2019; Cianci et al., 2024). Several analytical methods-such as the ΔCq method, Cq ratio, and direct quantification of transcript abundance-have been employed to model the relationship between mRNA degradation and PMI (Kimura et al., 2011; Ali et al., 2017; Alshehhi and Haddrill, 2019; Wang et al., 2021). However, mRNA stability is affected by multiple variables, including tissue type, temperature, humidity, and time since death (Bauer, 2007; Sampaio-Silva et al., 2013). Moreover, the suitability of specific postmortem marker genes differs among species, underscoring the need to identify honey bee-specific markers for PMI estimation.
In this context, the present study aimed to identify reliable molecular markers that can be used to estimate PMI in nurse bee (A. mellifera) carcasses. To this end, the expression levels of ten housekeeping genes [RPS5 (40S ribosomal protein S5), RPS18 (40S ribosomal protein S18), GAPDH (Glyceraldehyde-3-phosphate dehydrogenase), ARF1 (ADP-ribosylation factor 1), PPI (Peptidyl-prolyl cis-trans isomerase), PGK (Phosphoglycerate kinase), SDH (Succinate dehydrogenase flavoprotein subunit), TBP (TATA-box-binding protein), EF1 (Elongation factor 1-alpha F2), and EcR (Ecdysone receptor)] and four putative postmortem genes-LOC410857 (protein lethal (2) essential for life), DAAM (disheveled-associated activator of morphogenesis-like protein), and Prx2 (Peroxiredoxin 1)-were examined using quantitative real-time PCR (qRT-PCR). Expression ratios between candidate genes and stable reference genes were further analyzed to enhance resolution across PMI time points. Mathematical regression models were then established to determine the most suitable markers and to propose an accurate method for estimating PMI in honey bee carcasses.
MATERIALS AND METHODS
1. Sample preparation
Colonies of the western honey bee (A. mellifera) were maintained at the experimental apiary in Sangju-si, Korea. Nurse bees were collected based on their labor-specific behaviors (Johnson, 2010) and stabilized in cages (Bugdorm-1, MegaView Science Co., Ltd., Taichung City, Taiwan) supplied with 50% sucrose solution. The bees were maintained for 24 h in a dark incubator at 32℃ (JS Research Inc.) prior to experiments. To obtain carcasses for different postmortem intervals (PMIs), the nurse bees were euthanized with CO2 and maintained under controlled laboratory conditions. Carcasses were sampled at 1, 2, 3, 4, 5, 6, and 7 days postmortem, ensuring no fungal growth on the body surface. Samples were immediately stored at -80℃ until RNA extraction.
2. Candidate postmortem marker genes and primer design
Thirteen candidate postmortem marker genes were selected to evaluate their potential for PMI estimation: RPS5, RPS18, GAPDH, ARF1, PPI, PGK, SDH, TBP, EF1, LOC410857, DAAM, Prx2, and EcR gene was used as a reference gene based on its previously validated expression stability under postmortem conditions (Kim and Kim, 2025a). Primer sequences for ten genes (RPS5, RPS18, GAPDH, ARF1, RAB1a, PPI, PGK, SDH, TBP, and EF1) were obtained from previous reports (Moon et al., 2018; Jeon et al., 2020; Kim et al., 2022; Kim and Kim, 2025a, 2025b). Primers for LOC410857, DAAM, Prx2, and EcR were designed using the OligoCalc software (http://biotools.nubic.northwestern.edu/OligoCalc.html). PCR amplification efficiency was determined from standard curves derived from 10-fold serial dilutions of cDNA, and calculated using the formula E=10-1/slope. All primer sets exhibited efficiencies between 92-107% with coefficients of determination (R2)>0.99, indicating acceptable performance for qRT-PCR analysis(Table 1).
3. RNA extraction, cDNA synthesis, qRT-PCR
For each sample, the antennae were removed to prevent contamination from damaged tissue. Total RNA was extracted from individual heads using the yesR Total Plus Kit (GenesGen, Busan, Korea) according to the manufacturer’s protocol. Genomic DNA was eliminated using gDNA Eliminator Buffer and DNase I(GenesGen). The concentration and purity of RNA were measured using a SpectraMax QuickDrop spectrophotometer(Molecular Devices, Sunnyvale, CA, USA). RNA samples were stored at -80℃ until use.
Complementary DNA (cDNA) was synthesized from 50 ng of total RNA using ReverTra AceTM qPCR RT Master Mix (Toyobo, Japan) following the manufacturer’s protocol. The thermal cycling conditions were 37℃ for 15 min, 50℃ for 5 min, and 98℃ for 5 min.
qRT-PCR was performed on a CFX Connect Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA) using Thunderbird SYBR qPCR Master Mix (Toyobo, Osaka, Japan). The reaction conditions were as follows: initial denaturation at 95℃ for 1 min, followed by 40 cycles of 95℃ for 15 s, 56℃ for 15 s, and 72℃ for 30 s. Each reaction was run in triplicate. Amplification specificity was confirmed by melting curve analysis. Cq values were obtained automatically using the CFX Manager software, and relative expression levels were normalized to RAB1a using the 2-ΔΔCq method.
4. Statistical analysis and mathematical modeling
Normalized expression levels of each candidate gene across different PMIs were analyzed using one-way analysis of variance (ANOVA) followed by Tukey’s multiple comparison test (p<0.05). Expression ratios were calculated using the following formula:
Stable genes used as denominators were selected based on their expression stability rankings, determined by the smallest average standard deviation of expression levels across all PMIs (Fig. 1) and supported by a previous study (Kim and Kim, 2025a).
Cubic polynomial regression models were constructed to evaluate the correlation between gene expression (or expression ratios) and PMIs. The model with the highest coefficient of determination (R2) was considered the most accurate for PMI estimation.
RESULTS
1. Expression patterns of candidate postmortem marker genes
The expression levels of thirteen candidate genes were quantified in nurse bee heads at seven postmortem intervals (1-7 days) using qRT-PCR (Fig. 1). Among them, five genes-RPS5, RPS18, GAPDH, PPI, and EF1-exhibited significant temporal changes after death (p<0.05), whereas the remaining eight genes (ARF1, PGK, SDH, TBP, LOC410857, DAAM, Prx2, and EcR) showed no significant differences across PMIs (p>0.05).
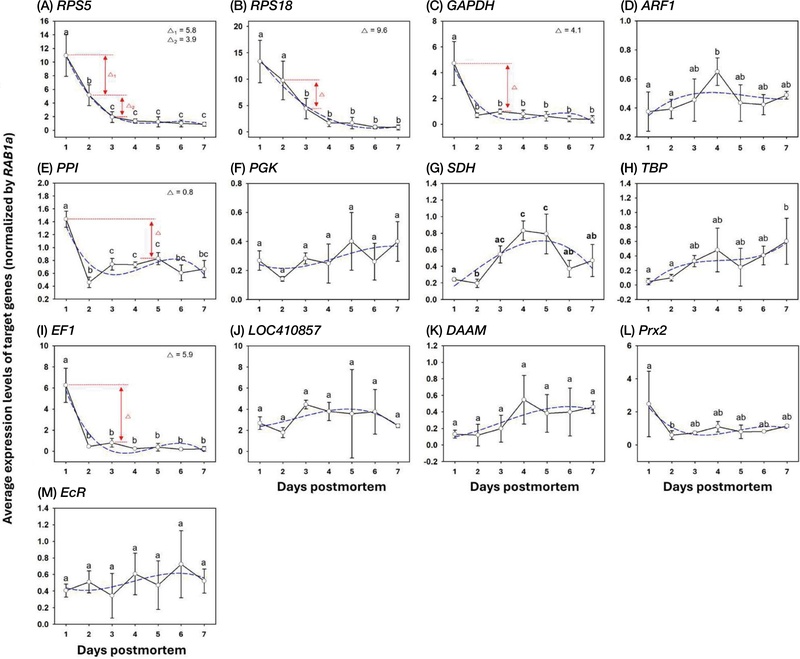
Expression patterns of postmortem marker genes under postmortem condition. The expression levels(average±standard deviation) of target genes were measured depending on PMI. The difference value (∆) represented the smallest gap between expression levels in PMI estimation ranges. The Different letters indicate significantly different values (p<0.05) that analyzing by One-way ANOVA with Turkey’s multiple comparison test.
Expression of RPS5 decreased markedly during the first 3 days postmortem, with statistically distinct levels at 1, 2, and 3 days (p<0.05). Thereafter, expression remained stable from 3 to 7 days, suggesting that RPS5 is a sensitive marker for distinguishing early (≤3 days) and late (≥3 days) postmortem phases (Fig. 1A). Similarly, RPS18 expression gradually declined from 1 to 4 days postmortem, with a clear separation between early (1-2 days) and mid-to-late (3-7 days) PMIs (p<0.05; Fig. 1B). In contrast, GAPDH, PPI, and EF1 showed significant downregulation only at 1 day postmortem (p<0.05), indicating limited utility for distinguishing longer PMIs (Fig. 1C-E).
Cubic regression analysis revealed strong correlations between PMI and the expression levels of RPS5 (R2=0.997) and RPS18 (R2=0.982), while other genes showed weaker associations (R2<0.90; Table 2). These results suggest that RPS5 and RPS18 can serve as quantitative indicators for estimating postmortem duration within 3 days after death.
2. Expression ratios of candidate genes
To improve PMI resolution, expression ratios were calculated between the five PMI-responsive genes (RPS5, RPS18, GAPDH, PPI, EF1) and the four stably expressed genes (ARF1, PPI, PGK, SDH). Among all combinations, ratios using SDH as the denominator exhibited the clearest differentiation among PMIs (Fig. 2). The RPS5/SDH and RPS18/SDH ratios significantly distinguished 1-, 2-, and 3-7-day postmortem groups (p<0.05), while GAPDH/SDH and EF1/SDH only separated 1-day samples from later time points. Ratios involving other denominators (e.g., RPS5/PPI, RPS18/PGK) showed partial separation between early (1-2 days) and late (3-7 days) intervals but exhibited lower model accuracy (R2<0.9).
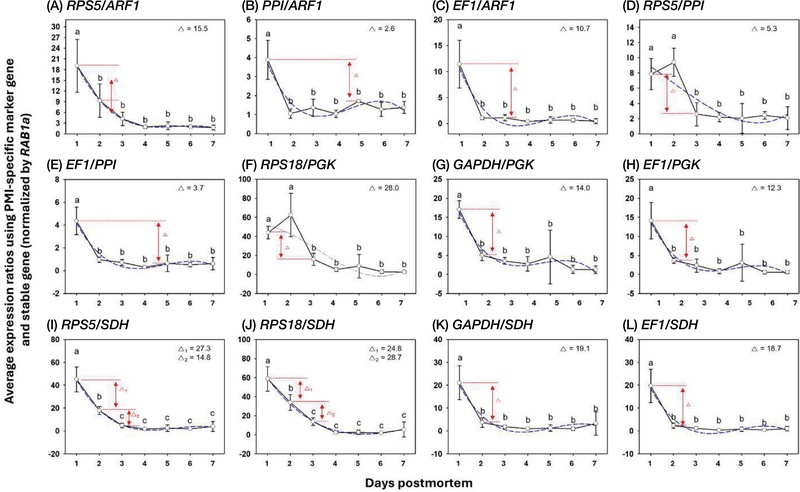
The ratios of expression levels using PMI-specific marker genes and stable gene under postmortem condition. The expression ratios (average±standard deviation) showing distinct PMI estimated ranges were calculated using PMI-specific gene and stably expressed gene (ARF1, PPI, PGK, and SDH). The difference value (∆) represented the smallest gap between expression levels in PMI estimation ranges. The Different letters indicate significantly different values (p<0.05) that analyzing by One-way ANOVA with Turkey’s multiple comparison test.
In contrast, the cubic regression models for RPS5/SDH and RPS18/SDH achieved high coefficients of determination (R2=0.996 for both; Table 2), indicating strong predictive relationships between expression ratios and PMIs.
The mean difference values between PMI groups were also greatest for these two ratios: 27.3 between 1 and 2 days for RPS5/SDH and 53.5 between 1 and 3-7 days for RPS18/SDH. These results demonstrate that both ratios can reliably distinguish among multiple postmortem stages and are suitable molecular markers for PMI estimation in A. mellifera.
3. Summary of optimal regression models
Cubic polynomial models were generated to describe the relationship between gene expression (or ratio) and PMI (Table 2). The models for RPS5, RPS18, RPS5/SDH, and RPS18/SDH exhibited R2>0.98, demonstrating high accuracy for PMI prediction. The regression equations are summarized in Table 2. These models indicate that PMI estimation based on expression ratios provides slightly higher precision than estimation based on single-gene expression levels.
Overall, the results demonstrate that (1) RPS5 and RPS18 show predictable degradation patterns after death; (2) ARF1, PPI, PGK, and SDH are stable postmortem reference genes; and (3) the ratios RPS5/SDH and RPS18/SDH provide the highest accuracy for PMI estimation in nurse bee carcasses, effectively distinguishing carcasses within 1-2 days postmortem from those after 3 days.
DISCUSSION
The present study aimed to identify candidate postmortem marker genes that can be used to estimate postmortem intervals (PMIs) in nurse bee (A. mellifera) carcasses. Estimating PMI in dead bees is crucial for understanding the causes of colony losses, since beekeepers usually detect mortality events after colony collapse has already occurred. Previous studies have primarily focused on the physiological effects of stressors on living bees (Zaobidna et al., 2017; Dolezal and Toth, 2018; Butolo et al., 2021; You et al., 2025), but few have explored molecular indicators in carcasses.
In this study, the expression levels of thirteen candidate genes were analyzed across seven postmortem intervals. Among them, RPS5 and RPS18 showed clear, time-dependent degradation patterns, allowing the differentiation of carcasses within the first three days after death. The cubic regression models of these genes exhibited strong correlation with PMI (R2=0.997 and 0.982, respectively), confirming their reliability as postmortem molecular markers. In contrast, other genes (GAPDH, PPI, and EF1) showed significant decreases only at 1 day postmortem, indicating limited applicability for PMI estimation beyond the early phase.
Interestingly, both RPS5 and RPS18 have been used as reference genes under various experimental conditions, including postmortem tissues in other species (Sampaio-Silva et al., 2013; Lv et al., 2017; Wang et al., 2022). However, the present study revealed that their expression levels were not stable after death in honey bee tissues, as confirmed by BestKeeper and geNorm analyses in our previous study (Kim and Kim, 2025a). This instability suggests that RPS5 and RPS18 are unsuitable as reference genes but rather are potential postmortem markers whose degradation correlates strongly with time since death. Such discrepancies with previous studies likely reflect interspecific differences in tissue physiology and mRNA stability, as well as methodological variations in sample storage and qRT-PCR conditions.
To improve PMI estimation accuracy, this study also analyzed expression ratios between dynamic and stable genes. According to the previous study (Kim and Kim, 2025a), four genes (ARF1, PPI, PGK, and SDH) used as denominator were ranked among the top four in Ref Finder, confirming their high expression stability under postmortem conditions. In this study, these genes also showed low standard deviations of expression levels across all PMIs, supporting their suitability as denominators for calculating expression ratios in PMI estimation. Among all tested combinations, the RPS5/SDH and RPS18/SDH ratios showed the highest discrimination power among postmortem groups and exhibited strong polynomial relationships with PMI(R2=0.996 for both). These ratios reliably distinguished carcasses at 1, 2, and ≥3 days postmortem, extending the estimation range and improving accuracy compared with single-gene models. This finding supports previous research in which the use of gene expression ratios was shown to reduce variability and improve model robustness(Kimura et al., 2011; Cianci et al., 2024).
The superior performance of RPS5/SDH and RPS18/SDH may be attributed to two factors: (1) the high stability of SDH expression postmortem, and (2) the consistent degradation kinetics of RPS5 and RPS18. Since ribosomal protein genes are highly expressed in active cells and rapidly degrade after cell death, their relative decrease provides a sensitive molecular signature of tissue decay. This suggests that ribosomal gene degradation can serve as a universal postmortem indicator across species.
Although the regression models developed in this study showed high accuracy under controlled laboratory conditions, several factors may influence their performance in the field. Temperature, humidity, and microbial activity can alter mRNA degradation rates (Bauer, 2007; Sampaio-Silva et al., 2013), potentially affecting PMI estimates. Therefore, field validation studies incorporating environmental variables are required to establish practical thresholds for PMI estimation in natural conditions.
Overall, this study provides the first molecular approach to PMI estimation in honey bees using gene expression dynamics. The findings demonstrate that RPS5 and RPS18, along with their ratios to SDH, can serve as reliable molecular markers to differentiate early and late postmortem phases in A. mellifera carcasses. This molecular framework may contribute to future forensic analyses of colony mortality events and improve diagnostic accuracy in apicultural investigations.
Acknowledgments
This study was supported by a research fund (RS-2025-02303308) from the Rural Development Administration, Republic of Korea. This manuscript is based on the master’s thesis of HeeJin Kim.
References
-
Ali, M. M., S. F. Ibrahim and A. A. Fayed. 2017. Using skin gene markers for estimating early postmortem interval at different temperatures. Am. J. Forensic Med. Pathol. 38: 323-325.
[https://doi.org/10.1097/PAF.0000000000000337]

-
Alshehhi, S. and P. R. Haddrill. 2019. Estimating time since deposition using quantification of RNA degradation in body fluid-specific markers. Forensic Sci. Int. 298: 58-63.
[https://doi.org/10.1016/j.forsciint.2019.02.046]

-
Bauer, M. 2007. RNA in forensic science. Forensic Sci. Int. Genet. 1: 69-74.
[https://doi.org/10.1016/j.fsigen.2006.11.002]

-
Brooks, J. W. and L. Sutton. 2017. Postmortem changes and estimating the postmortem interval. In Veterinary Forensic Pathology, Volume 1 (pp. 43-63). Springer.
[https://doi.org/10.1007/978-3-319-67172-7_4]

-
Butolo, N., P. Azevedo, L. Alencar, O. Malaspina and R. Nocelli. 2021. Impact of low temperatures on the immune system of honeybees. J. Therm. Biol. 101: 103082.
[https://doi.org/10.1016/j.jtherbio.2021.103082]

-
Cianci, V., C. Mondello, D. Sapienza, M. C. Guerrera, A. Cianci, A. Cracò, F. Omero, V. Gioffrè, P. Gualniera and A. Asmundo. 2024. Potential Role of mRNA in Estimating Postmortem Interval: A Systematic Review. Int. J. Mol. Sci. 25: 8185.
[https://doi.org/10.3390/ijms25158185]

-
Dolezal, A. G. and A. L. Toth. 2018. Feedbacks between nutrition and disease in honey bee health. Curr. Opin. Insect Sci. 26: 114-119.
[https://doi.org/10.1016/j.cois.2018.02.006]

- IPBES. 2016. The Assessment Report of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services on Pollinators, Pollination and Food Production Bonn, Germany.
-
Jeon, J. H., K. Moon, Y. Kim and Y. H. Kim. 2020. Reference gene selection for qRT-PCR analysis of season- and tissue-specific gene expression profiles in the honey bee Apis mellifera. Sci. Rep. 10: 13935.
[https://doi.org/10.1038/s41598-020-70965-4]

-
Johnson, B. R. 2010. Division of labor in honeybees: form, function, and proximate mechanisms. Behav. Ecol. Sociobiol. 64: 305-316.
[https://doi.org/10.1007/s00265-009-0874-7]

- Jung, C. 2008. Economic value of honeybee pollination on major fruit and vegetable crops in Korea. J. Apic. 23: 147-152.
-
Kim, H. and Y. H. Kim. 2025a. Selection of Stable Reference Genes for qRT-PCR in Nurse Bee (Apis mellifera) under Postmortem Conditions. J. Apic. 40: 211-224.
[https://doi.org/10.17519/apiculture.2025.09.40.3.211]

-
Kim, H. and Y. H. Kim. 2025b. Evaluation of Reference Genes for Quantitative Real-Time PCR in Honey Bee, Apis mellifera, Under Various Miticide Exposure Conditions. Entomol. Res. 55: e70026.
[https://doi.org/10.1111/1748-5967.70026]

-
Kim, H.-K. 2022. The Effect of Honey Bee Mites on the Winter Colony Loss. J. Apic. 37: 291-299.
[https://doi.org/10.17519/apiculture.2022.09.37.3.291]

-
Kim, S., S. Cho and S. H. Lee. 2022. Selection of stable reference genes for real-time quantitative PCR in honey bee pesticide toxicity studies. J. Apic. Res. 61: 26-36.
[https://doi.org/10.1080/00218839.2021.1950972]

-
Kimura, A., Y. Ishida, T. Hayashi, M. Nosaka and T. Kondo. 2011. Estimating time of death based on the biological clock. Int. J. Legal Med. 125: 385-391.
[https://doi.org/10.1007/s00414-010-0527-4]

-
Lee, S.-J., S.-H. Kim, J. Lee, J.-H. Kang, S.-M. Lee, H. J. Park, J. Nam and C. Jung. 2022. Impact of Ambient Temperature Variability on the Overwintering Failure of Honeybees in South Kore. J. Apic. 37: 331-347.
[https://doi.org/10.17519/apiculture.2022.09.37.3.331]

-
Lv, Y.-H., J.-L. Ma, H. Pan, Y. Zeng, L. Tao, H. Zhang, W.-C. Li, K.-J. Ma and L. Chen. 2017. Estimation of the human postmortem interval using an established rat mathematical model and multi-RNA markers. Forensic Sci. Med. Pathol. 13: 20-27.
[https://doi.org/10.1007/s12024-016-9827-4]

-
Meixner, M. D. 2010. A historical review of managed honey bee populations in Europe and the United States and the factors that may affect them. J. Invertebr. Pathol. 103: S80-S95.
[https://doi.org/10.1016/j.jip.2009.06.011]

-
Moon, K., S. H. Lee and Y. H. Kim. 2018. Validation of quantitative real-time PCR reference genes for the determination of seasonal and labor-specific gene expression profiles in the head of Western honey bee, Apis mellifera. PLoS One. 13: e0200369.
[https://doi.org/10.1371/journal.pone.0200369]

-
Pozhitkov, A. E., R. Neme, T. Domazet-Lošo, B. G. Leroux, S. Soni, D. Tautz and P. A. Noble. 2017. Tracing the dynamics of gene transcripts after organismal death. Open Biol. 7: 160267.
[https://doi.org/10.1098/rsob.160267]

-
Reilly, J., D. Artz, D. Biddinger, K. Bobiwash, N. Boyle, C. Brittain, J. Brokaw, J. Campbell, J. Daniels and E. Elle. 2020. Crop production in the USA is frequently limited by a lack of pollinators. Proc. R. Soc. B 287: 20200922.
[https://doi.org/10.1098/rspb.2020.0922]

-
Sampaio-Silva, F., T. Magalhaes, F. Carvalho, R. J. Dinis-Oliveira and R. Silvestre. 2013. Profiling of RNA degradation for estimation of post morterm interval. PLoS One. 8: e56507.
[https://doi.org/10.1371/journal.pone.0056507]

-
Scrivano, S., M. Sanavio, P. Tozzo and L. Caenazzo. 2019. Analysis of RNA in the estimation of post-mortem interval: a review of current evidence. Int. J. Legal Med. 133: 1629-1640.
[https://doi.org/10.1007/s00414-019-02125-x]

-
Shafeeq, T., A. Malik, A. Zaheer and K.-Y. Lee. 2020. Differentiated transcriptional regulation before and after death in Indianmeal moth Plodia interpunctella. J. Asia. Pac. Entomol. 23: 1096-1100.
[https://doi.org/10.1016/j.aspen.2020.09.009]

- Sumner, D. A. and H. Boriss. 2006. Bee-conomics and the leap in pollination fees. Agric. Resour. Econ. 9: 9-11.
-
Wang, Y., M. Wang, C. Luo, L. Li, W. Xu, G. Hu, Y. Wang, J. Amendt and J. Wang. 2021. Dynamics of insects, microorganisms and muscle mRNA on pig carcasses and their significances in estimating PMI. Forensic Sci. Int. 329: 111090.
[https://doi.org/10.1016/j.forsciint.2021.111090]

-
Wang, Y., M. Wang, W. Xu, Y. Wang, Y. Zhang and J. Wang. 2022. Estimating the postmortem interval of carcasses in the water using the carrion insect, brain tissue RNA, bacterial biofilm, and algae. Front. Microbiol. 12: 774276.
[https://doi.org/10.3389/fmicb.2021.774276]

-
Wenzlow, N., D. Mills, J. Byrd, M. Warren and M. T. Long. 2023. Review of the current and potential use of biological and molecular methods for the estimation of the postmortem interval in animals and humans. J. Vet. Diagn. Invest. 35: 97-108.
[https://doi.org/10.1177/10406387231153930]

-
You, E., J. Cha, H. Kim and Y. H. Kim. 2025. Comparison of the toxicity and potential ecological risks of various pesticides for nurses of honey bee (Apis mellifera. L). J. Environ. Chem. Ecotoxicol. 7: 791-801.
[https://doi.org/10.1016/j.enceco.2025.04.008]

-
Zaobidna, E. A., K. Żółtowska and E. Łopieńska-Biernat. 2017. Varroa destructor induces changes in the expression of immunity-related genes during the development of Apis mellifera worker and drone broods. Acta Parasitol. 62: 779-789.
[https://doi.org/10.1515/ap-2017-0094]

