
Thermal Stress-related Genes in Honey Bees (Apis mellifera Linnaeus) by RNA-seq Analysis
Abstract
The honey bee, Apis mellifera, a vital pollinator for agricultural crops, has experienced a decline in numbers over the past decades due to colony collapse disorder (CCD). The suspected causes of CCD have included climate change, parasites, and pesticide use. This study focused on the impact of climate change, as insect body temperature, which is regulated by environmental conditions, is highly susceptible to thermal stress. We investigated the molecular response processes of A. mellifera by comparing gene expression profiles under low (10°C) and high (38°C) temperature stress with those of an untreated control (25°C). RNA-seq analysis identified 349,489 assembled transcripts. Differentially expressed genes (DEGs) with a log2 fold change ≥ 2 were compared between the temperature conditions. We identified 2,838DEGs at 10°C and 3,634DEGs at 38°C. Specifically, 2,077 and 2,007 genes were upregulated, while 1,438 and 3,393 genes were downregulated at 10°C and 38°C, respectively, compared to 25°C. The honey bees exhibited increased expression of HSP70 and HSP60 genes under high-temperature stress, whereas ApoID, HR38, and ABRA genes were highly expressed under low-temperature stress. ETH, TTLL4, and OR genes were highly expressed under both temperature conditions. Our findings demonstrate that honey bees exhibit distinct transcriptomic responses to thermal stress, with HSPs mediating heat tolerance and calcium/lipid-associated genes supporting cold tolerance.
Keywords:
Apis mellifera, Climate change, Colony collapse disorder, DEG, Thermal stress, PollinatorINTRODUCTION
Honey bees (Apis mellifera Linnaeus) play a crucial role in pollinating the nectar of various crops and flowers, with approximately one-third of crops relying on honey bee pollination (Gallai et al., 2009; Hou et al., 2014). Also, honey bees have economical effect estimated about US$200-500 billion per annual (Lautenbach et al., 2012). Colony collapse disorder (CCD), which is responsible for destroying colonies of honey bees, has been reported as a global concern. For example, the number of honey bee colonies in the USA declined from 4.6 million in 1970 to 2.7 million in 2014, in Europe from 21.1 million in 2014 and from 21.1 million to 17.7 million during the same period (FAO, 2017; Paris et al., 2017). More recently, significant colony losses have been reported in South Korea. According to Lee et al. (2022), approximately 43% of bee hives in Jeollanam-do were affected, resulting in the loss of about 105,900 colonies. In Gyeongsangnam-do, 11.1% of hives were lost, totaling about 38,400 colonies, while in Jeju, 15.1% of bees disappeared from 74,200 colonies, affecting 31.3% of beekeeping farms. The suspected causes of CCD include global climate change, pesticide exposure, nutritional deficiencies, and pathogens (vanEngelsdorp et al., 2009; Baines et al., 2017; Mahanhuda and Tiwari, 2024).
Global climate change gives many negative effects on honey bees, particularly because they are ectotherms which rely on environmental heat sources for their body temperature (Hilderbrand and Goslow Jr., 2001). When global warming gives thermal stress to honey bees, heat shock protein (HSP) family do key play role on heat stress tolerance. HSPs play role of heat hardening in high temperature, which in turn provides protection from subsequent and more severe temperature (Matz et al., 1995). HSP includes large HSPs, HSP90, HSP70, HSP60, HSP40, and small HSPs. Many HSPs perform chaperone functions by stabilizing new proteins to promote correct folding or by helping to refold proteins that were damaged by cellular stress (De Maio, 1999). HSP family is essential for relieving physiological stress(Ceylan et al., 2023). To regulate the expression of HSPs, heat shock factors (HSFs) are considered transcription factors of HSPs within the cellular system (Watanabe and Tanaka, 2018). In Caenorhabditis elegans, HSF regulates approximately 942 genes during heat stress for homeostasis beyond the immediate stress response (Brunquell et al., 2016). HSP90 gene participates the protein conformational maturation (Hu et al., 2022). HSP70 gene is conserved molecular chaperon ubiquitously expressed in prokaryote and eukaryote organism (Rosenzweig et al., 2019). Seventeen kinds of HSP70 genes are encoded in human (Brocchieri et al., 2008). HSP60 gene is divided into two groups of chaperonins which are expressed in different locations within cells. HSP60 is categorized into 2 groups of chaperonins located in different parts of the cell. Group I of HSP60s are found in prokaryotes, as well as in the mitochondria and chloroplasts of eukaryotes. Group II of HSP60s are present in archaea and the cytosol of eukaryotic cells(Balchin et al., 2018). Octopamine coordinates thermo-protection of an insect nervous system(Armstrong and Robertson, 2006).
Despite climate change resulting in colder winter, insects have developed seasonal adaptations to cold stress, including environmentally programmed periods of dormancy called diapause (Denlinger et al., 2005; Kostál, 2006), and rapid cold hardening (RCH), which provides cold resistance against rapidly decreased temperatures (Lee Jr. et al., 1987). A remarkable feature of RCH is that cells directly respond to low temperature in the absence of stimulation from the brain and hormones. Although the mechanism and down-stream of cold-hardening pathway are unknown (Yi and Lee Jr., 2004; Teets et al., 2008), chilling causes a rapid melanogaster, a mutation of membrane protein, dystroglycan, have elevated intracellular calcium levels, which correlates with improved performance and survival at low temperatures. To regulate localization of calcium in rapid cold condition, calcium and calmodulin-dependent signaling event have important roles(Teets et al., 2013).
In this study, we investigated the transcriptional responses of honey bees to thermal stress. Transcriptome analysis identified differentially expressed genes(DEGs), and selected candidates were validated using quantitative real-time PCR (RT-qPCR). Our results demonstrate that honey bees exhibit distinct transcriptomic responses to thermal stress, with heat shock proteins mediating heat tolerance and calcium/lipid-associated genes supporting cold tolerance.
MATERIALS AND METHODS
1. Honey bee rearing and thermal stress treatment
The honey bee apiaries were maintained at the experimental apiary of Gyeongkuk National University. Worker bees were collected using an insect net for experiment and placed in insect breeding cages(120×80mm)(SPL, Pocheon, Korea) to stabilize honey bees in a new environment (Ulziibayar et al., 2025). Collected bees were transferred to the laboratory and maintained at 25±2℃ with 50-60% of relative humidity, and provided with a 40% sugar solution as food for one day. To expose the honey bees to thermal stress conditions, they were kept at 28℃, 10℃, and 38℃ for 12 h. Following these treatments, the honey bees were used for RNA extraction.
2. RNA extraction and cDNA library construction
Total RNA from A. mellifera was isolated using Trizol reagent (Invitrogen, Carlsbad, CA, USA) according to manufacturer’s instructions. To generate cDNA library, TrueSeq stranded total RNA with Ribo-Zero H/M/R_Gold (Illumina, California, USA) was used. The sequencing was performed on a Novaseq (Illumina, California, USA) platform by Macrogen (Seoul, Korea). The sequences were trimmed using Trimmomaic 0.38. The trimmed sequences were assembled by Trinity (version trinityseq_r20140717, bowite 1.1.2) via de novo sequencing. Total reads were mapped using RSEM version 1.3.1, bowtie 1.1.2 from http://genontology.org/, and BLAST version 2.9+ (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and DIAMOND version 0.9.21 (https://github.com/bbuchfink/diamond/releases?after=v0.9.21) were used for gene annotation. For DEG analysis, size factors were estimated from count data and normalized using the trimmed mean of M-values(TMM) method in edgeR library in R (Bioconductor, Seattle, WA, USA).
3. In silico analysis
The mRNA sequencing was performed using the Nova Seq platform. The sequenced reads were processed to remove adapter sequences using Trimmomatic 0.38. Subsequently, de novo assembly of the RNA sequences was conducted using Trinity (version trinityseq_r20140717, bowtie 1.1.2). Gene expression levels were quantified by mapping the de novo assembled RNA sequences with RSEM (version v1.3.1, bowtie 1.1.2). Gene identification was performed through the DIAMOND, using databases obtained from the Gene Ontology (GO) knowledgebase (http://geneontology.org/) and the BLAST. The mapped data were then analyzed for differentially expressed genes(DEGs) using the edgeR library. The Fragmented per Kilobase of transcript per Million mapped reads(FPKM) values were utilized for DEG analysis.
4. Quantitative real time polymerase chain reaction(RT-qPCR)
cDNA was synthesized by Max RT premix kit(Intron, Seongnam, Korea) with random primer for RT-qPCR. The RNA was dissolved in diethyl pyrocarbonate (DEPC) treated water, and 500ng of this RNA was used for cDNA synthesis in a total volume of 20 μL. Reverse transcription and annealing were conducted at 45℃ for 60 min. Subsequently, inactivation of reverse transcriptase was performed at 95℃ for 5min. RT-qPCR was performed cDNA with iSYBR green premix (Bio-Rad) with the following conditions: initial denaturing at 95℃ for 3 min, followed by 40 cycles of 95℃ denaturing for 30 s, 50℃ annealing for 30 s, 72℃ extension for 30 s, and melting step. Each PCR reaction mixture (20 μL) contains 10 μL of iQ SYBR Green Supermix, 1 μL of each 10 pmol forward and reverse primer set (Table 1), 2 μL of cDNA (200 ng), and 6 μL of water. RNA expression levels were normalized following Comparative Ct (ΔΔCT) method (Livak and Schmittgen, 2001) against EF-1 as reference gene.
5. Graphics and statistical analysis
The heatmaps were demonstrated by R with heatmap library. Statistical values in the graphs are presented as mean±standard deviation. The results were visualized using GraphPad Prism 9 (Dotmatics, Boston, USA). One-way ANOVA was performed to analyze the means with the SAS program(SAS Institute Inc., 1989). Differences among treatment means were assessed using the LSD test, with a significance level set at P<0.05.
RESULTS
1. Abundance of transcriptome
The transcriptome of honey bees was analyzed and generated about 12.0 GB of sequences from each treatment(Fig. 1). 100 Mb pairs of reads were retrieved after trimming (Fig. 1A). After sequences assembly, about 73.68-87.00% sequences were mapped to genes. Total 349,489 assembled transcripts were annotated through DIAMOND version 0.9.21. Principal component analysis(PCA) was performed to compare honey bees exposed to high (38℃), low (10℃), and normal (control, 25℃) temperatures(Fig. 1B).
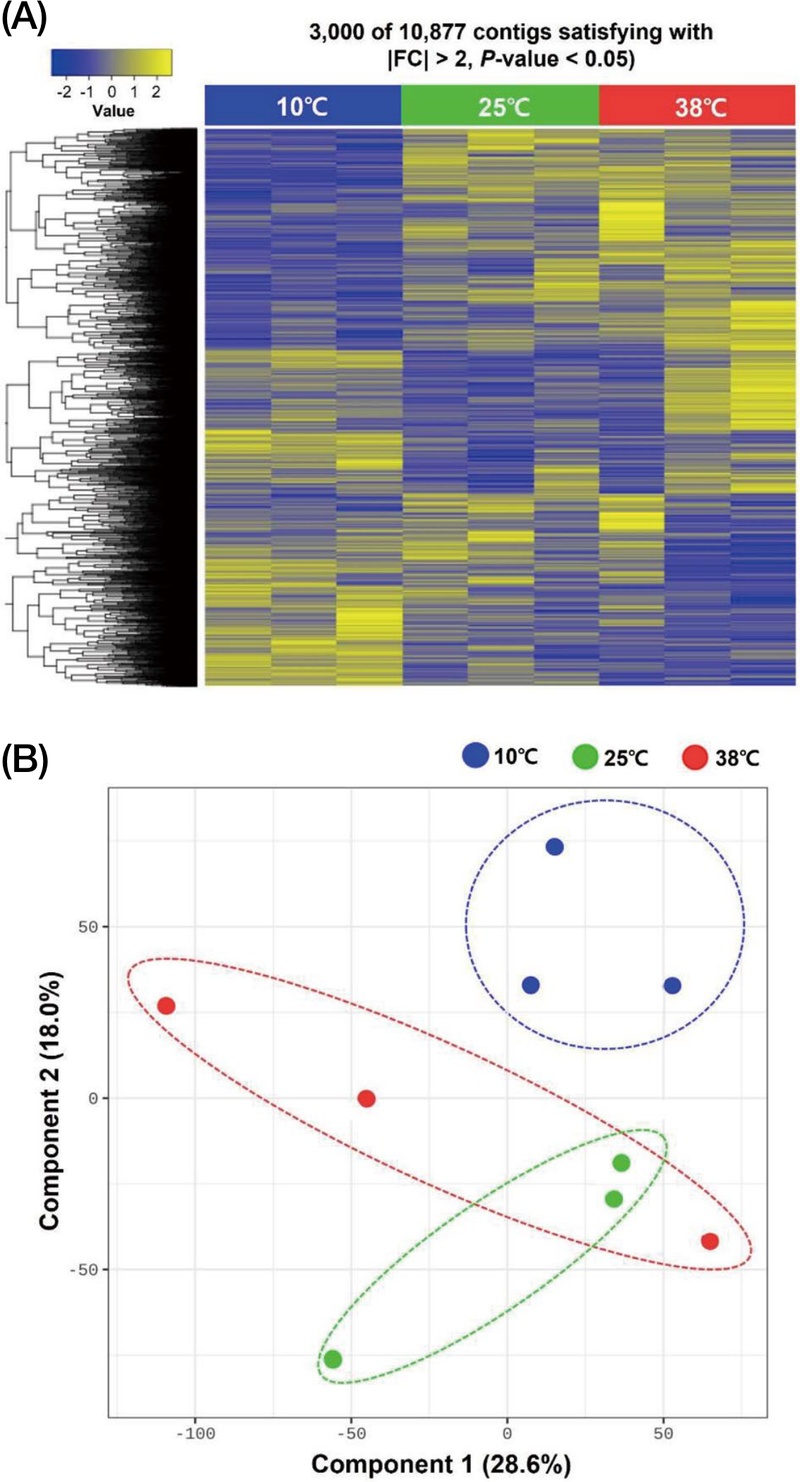
Heatmap and multidimentional scaling (MDS) analysis of differentially expressed genes in Apis mellifera among 3 different temperatures exposed workers. (A) The heatmap was generated using Z-scores transformed from Fragments per Kilobase of transcript per Million mapped reads values. (B) The MDS plot was generated using R programming language.
2. Analysis of differentially expressed genes(DEGs)
Transcriptional abundance of 38℃ and 10℃ were compared to control (25℃) via RSEM by using FPKM to perform DEG(Fig. 2). DEGs were selected based on a threshold of a 2-fold change (FC) value and a P-value of less than 0.05. Fig. 2A showed numbers of up- and down-regulated genes. Up- and down-regulated genes were generated 2,077 and 1,438 at 10℃ compared to 25℃, respectively (Fig. 2B and 2C). At 38℃ compared to 10℃, a total of 2,608 genes were up-regulated and 3,303 were down-regulated, whereas at 38℃ compared to 25℃, 2,077 genes were up-regulated and 3,393 were down-regulated (Fig. 2B and 2C). Total DEGs were performed GO term analysis followed by biological process, cellular components, molecular function. GO term analysis was displayed via bubble plot (Fig. 3). In case of biological process, GO:0008152 metabolic process showed the highest weighted score (139.06). GO:0009987 cellular process showed the highest number of count(900). In case of cellular component, GO:0044464 cell part showed the highest weighted score (156.61) and count (1007). In case of molecular function, GO:0005488 binding activity showed the highest weighted score (174.64) and count(651).
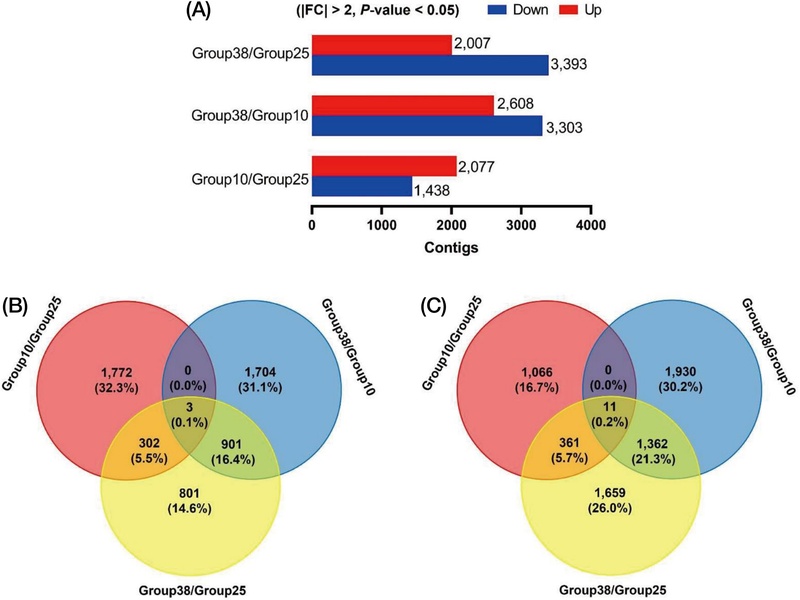
Analysis of differentially expressed genes(DEGs) in Apis mellifera among 3 different temperatures exposed workers. (A) DEGs abundance of A. mellifera among different temperature group. (B) Venn diagram analysis. The number of specific and shared DEGs were showed between three different temperature groups. All data were selected from threshold levels of >2-folds change and P-value<0.05 in Fragments per Kilobase of transcript per Million mapped reads values.
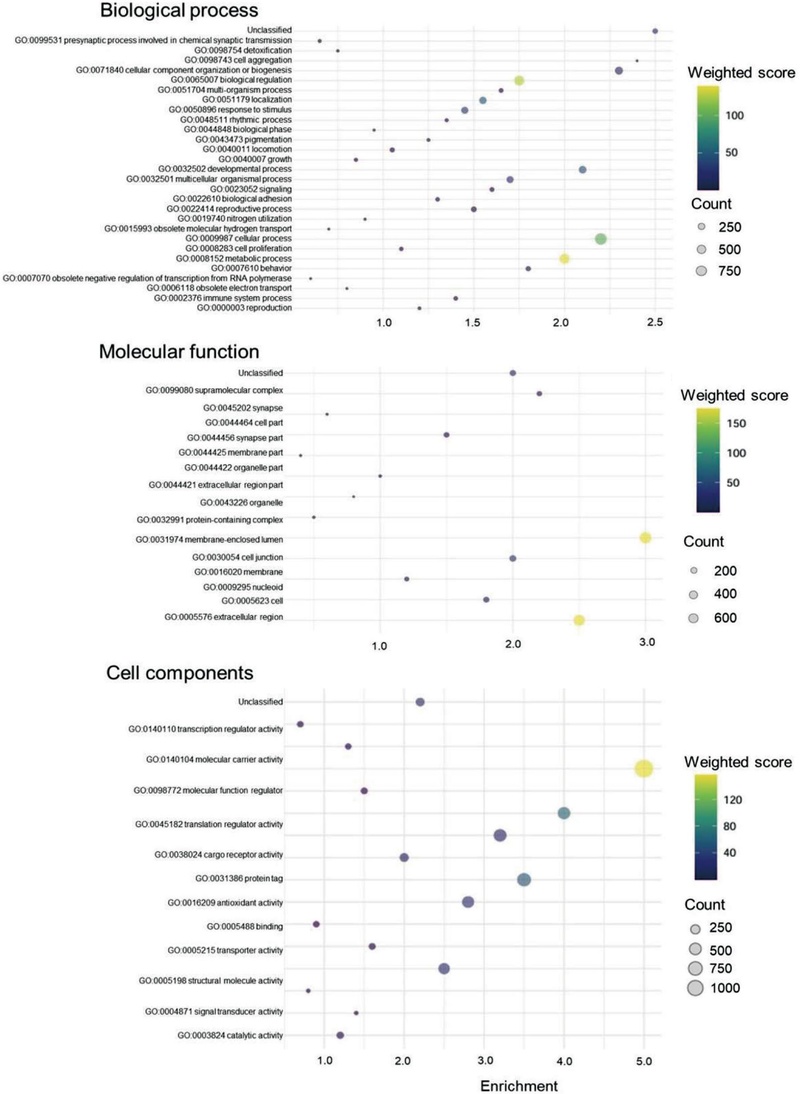
Gene Ontology (GO) classification of all annotated unigenes. 10,877 unigenes were assigned to three main GO categories of ‘Biological process with 29 functional subcategories’, ‘Molecular function with 16 functional subcategories’, and ‘Cellular component with 13 functional subcategories. Some unigenes were not classified and were shown as ‘Unclassified’ in each category.
3. Thermal stress related genes
HSP and cold stress responsible genes were selected from DEGs to analyze thermal stress responsible genes (Fig. 4). In high temperature exposed group, HSP70 (c109245_g1_i1), HSP90 (c150852_g2_i1), HSP83 (c107800_g1_i1), HSP60 (c24476_g1_i1) were highly expressed compared to normal temperature exposed group (Fig. 4A). These HSPs showed more than 2 FC value. In low temperature exposed group, calcium channel related proteins and calcium binding proteins were analyzed by heat map with FC values. Some genes were selected what is more than 2 FC value, Apolipolin D, Odorant receptor (OR), Hormone receptor 38 (HR38), Ecdysis trigger hormone (ETH), Tubulin tyrosine ligase like 4 (TTLL), Voltage dependent L-type calcium channel (VDLCC), and Actin binding Rho-activating (ABRA) from DEGs. RT-qPCR was conducted to validate the expression of genes associated with thermal stress(Fig. 5). In the group exposed to 38℃, HSP60, HSP70, OROa, ETH, and TTLL4a exhibited significantly different expression levels compared to the control as 25℃ (P<0.05; Fig. 5A). In the group exposed to 10℃, TTLL4b, ApolD, VDLCC, ABRA, and HR38 also showed significantly altered expression relative to the normal temperature as control(Fig. 5B).
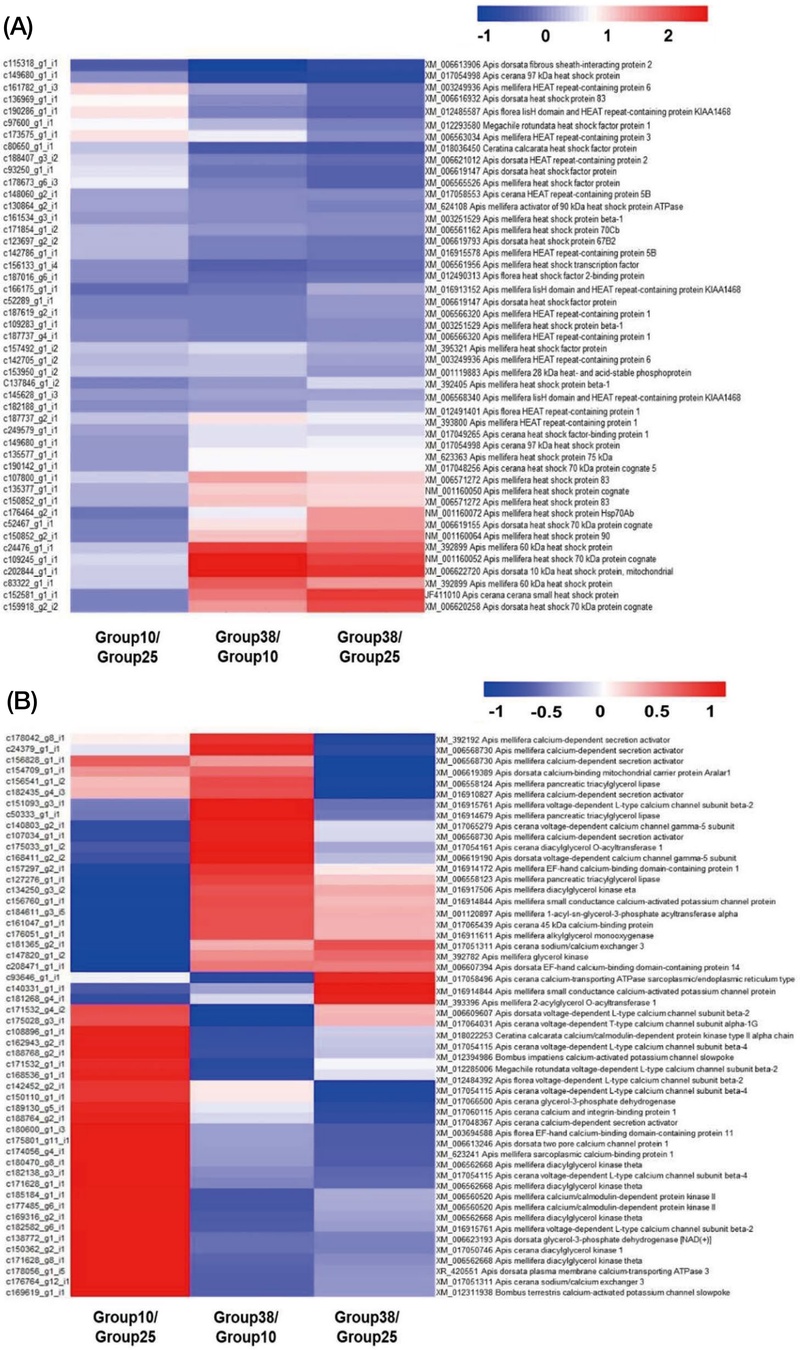
The heatmap analysis of heat shock protein and calcium related proteins in Apis mellifera among 3 different temperatures exposed workers. (A) Heat shock proteins from differentially expressed genes(DEGs) for high temperature tolerance. (B) Calcium related proteins from DEGs for low temperature tolerance.
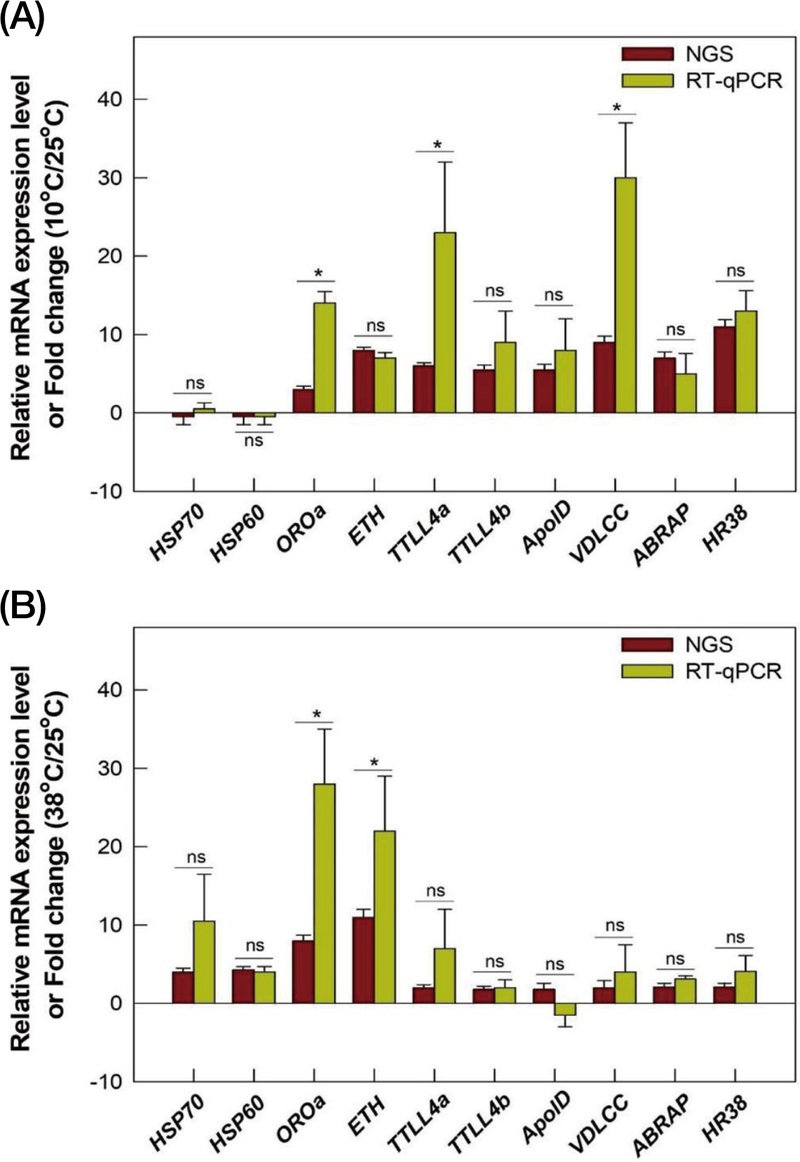
Validation of differentially expressed genes (DEGs) associated with thermal stress in Apis mellifera by RT-qPCR. (A) Gene expression of honeybee when honey bee got heat stress. (B) Gene expression of honeybee when honeybee got cold stress. All the samples were replicated independently three times. HSP70 and 60; Heat shock protein 70 and 60, OROa; Octopamine receptor_Oa, ETH; Ecdysis triggering hormone, TTLL4a and b; tubulin polyglutamylase TTLL4a and b, VDLCC; Voltage dependent L-type calcium channels, ABRAP; Actin binding rho activating protein, HR38; hormone receptor 38. Asterisks above standard deviation bars indicate significant difference between groups at Type I error=0.05 (LSD test). The letter “ns” stands for a non-significant difference between groups.
High-temperature exposure markedly upregulated genes related to proteolysis (trypsin-1), neurotransmission (octopamine receptor, GABA transporter), and stress adaptation (ecdysis triggering hormone, chemosensory proteins), with the highest FC observed for Aphis dorsata trypsin-1 in Table 3, while in low-temperature exposure, genes involved in developmental regulation (nuclear hormone receptor HR38, distal-less), muscle function (troponin H/I, muscle LIM protein 1), and cuticle formation (pupal cuticle protein-like, laccase 2) were predominantly upregulated, with the largest change detected for A. mellifera HR38 in Table 4.

Gene list of differentially expressed genes with highly altered Fragments per Kilobase of transcript per Million mapped reads (FPKM) values under high-temperature condition
DISCUSSION
Insects are the most diverse and successful group of animals, playing indispensable roles in terrestrial ecosystems (Foottit and Adler, 2017). Their life histories are strongly influenced by abiotic factors, among which temperature is particularly critical. Temperature governs insect population abundance and distribution (Li et al., 2020) and affects population dynamics and geographic ranges by altering developmental rates, life cycles, and the availability of host plants (Bale et al., 2002). Under global warming, more frequent extreme thermal events can directly and indirectly impact insect survival and disrupt a wide array of physiological and metabolic functions, including feeding, digestion, detoxification, reproduction, growth, and development (Garrad et al., 2016).
Previous studies have demonstrated that thermal stress elicits distinct transcriptional responses depending on whether insects are exposed to high- or low-temperatures (Vatanparast and Park, 2021, 2022; Vatanparast et al., 2021). Metabolizable energy is essential for maintaining homeostasis and supporting growth (Richard, 1986). Exposure to temperatures above the optimal range impairs both energy reserves and metabolic process(Belhadj et al., 2015). Under heat stress, the synthesis of most proteins decline, including ATPases involved in major metabolic pathways such as glycolysis, the tricarboxylic acid cycle, and oxidative phosphorylation (Li et al., 2020).
In our study, high-temperature exposure prominently induced genes related to proteolysis, neuronal modulation, and general stress adaptation, whereas low-temperature exposure favored genes involved in developmental regulation, muscle function, and cuticle remodeling. These patterns are consistent with established principles of insect thermal physiology: heat often drives rapid metabolic turnover and neuromodulation, while cold necessitates developmental, structural, and contractile adjustments to maintain function (Li et al., 2020).
HSP family plays a key role in protecting cellular proteins under high-temperature conditions. Elevated temperatures can disrupt stabilizing disulfide bonds, leading to protein denaturation and loss of function (Matz et al., 1995). Consistent with this, honey bees subjected to heat stress in our study exhibited significantly upregulated expression of HSP70 and HSP80, indicating activation of molecular chaperone systems to mitigate heat-induced protein damage (Fig. 5).
Interestingly, ETH (c15958_g1_i1), OROa (c181508_g1_i4), and TTLL4 (c182436_g1_i1) were upregulated under both high- and low-temperature conditions. In Drosophila, ETH expression increases under high temperatures to promote reproduction (Meiselman et al., 2017), and in the western black spider, 20E is elevated under high-temperature conditions (Moen et al., 2022). Our results suggest that elevated 20E levels may stimulate ETH expression under heat stress. However, ETH was also highly expressed under cold conditions, indicating a potential role in promoting survival under stressful environments, which warrants further investigation. TTLL4 accumulation in organelles can induce hyperglutamylation-related neurodegeneration, as observed in mice, suggesting a negative response to harsh conditions (Bodakuntla et al., 2021).
Cold tolerance in insects often relies on lipid accumulation, which can lower the freezing point of cells and protect against winter-induced damage (Trenti et al., 2022). In honey bees, apolipoprotein D expression increases under cold stress, potentially influencing hemolymph viscosity and circulation. Similarly, Abra plays an important role in supporting blood flow in human (Troidl et al., 2009). HR38 has been shown to regulate carbohydrate metabolism during mosquito reproduction (Dong et al., 2018).
Overall, extreme temperatures induce dynamic transcriptomic changes in honey bees. Under heat stress, HSPs facilitate protein folding and confer heat tolerance, whereas under cold stress, calcium signaling and lipid metabolism contribute to reducing cellular freezing points and enhancing cold tolerance. TTLL4 appears to mediate negative effects under harsh conditions. This transcriptomic analysis provides fundamental insights into the physiological mechanisms that enable honey bees to withstand extreme temperatures.
CCD has caused widespread damage to honey bee populations worldwide, yet its precise causes remain unclear. Climate change, pesticide overuse, and honey bee pests have been suggested as potential contributing factors. In this study, we investigated the comparative transcriptomic responses of honey bees under different temperature conditions to elucidate the effects of thermal stress, which is closely related to climate change. Collectively, these findings offer a valuable foundation for understanding honey bee thermal adaptation and have important implications for mitigating CCD.
Acknowledgments
This research was supported by a Research Grant of Gyeongkuk National University to YP.
References
-
Armstrong, G. A. B. and R. M. Robertson. 2006. A role for octopamine in coordinating thermo protection of an insect nervous system. J. Therm. Biol. 31(1-2): 149-158.
[https://doi.org/10.1016/j.jtherbio.2005.11.022]

-
Baines, D., E. Wilton, A. Pawluk, d. G. Michael and N. Chomistek. 2017. Neonicotinoids act like endocrine disrupting chemicals in newly-emerged bees and winter bee. Sci. Rep. 7: 10979.
[https://doi.org/10.1038/s41598-017-10489-6]

-
Balchin, D., G. Milicic, M. Strauss, M. Hayer-Hartl and F. U. Hartl. 2018. Pathway of actin folding directed by the eukaryotic chaperonin TRiC. Cell 174(6): 1507-1521.
[https://doi.org/10.1016/j.cell.2018.07.006]

-
Bale, J. S., G. J. Masters, I. D. Hodkinson, C. Awmack, T. M. Bezemer and V. K. Brown. 2002. Herbivory in global climate change research: direct effects of rising temperature on insect herbivores. Glob. Change Biol. Bioenergy 8: 1-16.
[https://doi.org/10.1046/j.1365-2486.2002.00451.x]

-
Belhadj, S. I., T. Najar, A. Ghram and M. Abdrrabba. 2015. Heat stress effects on livestock: molecular, cellular and metabolic aspects, a review. J. Anim. Physiol. Anim. Nutr. 100: 401-412.
[https://doi.org/10.1111/jpn.12379]

-
Bodakuntla, S., C. Janke and M. M. Magiera. 2021. Tubulin polyglutamylation, a regulator of microtubule functions, can cause neurodegeneration. Neurosci. Lett. 746: 135656.
[https://doi.org/10.1016/j.neulet.2021.135656]

-
Brocchieri, L., d. M. Conway, E. Macario and A. J. Macario. 2008. Hsp70 genes in the human genome: conservation and differentiation patterns predict a wide array of overlapping and specialized functions. BMC Evol. Biol. 8: 19.
[https://doi.org/10.1186/1471-2148-8-19]

-
Brunquell, J., S. Morris, Y. Lu, F. Cheng and S. D. Westerheide. 2016. The genome-wide role of HSF-1 in the regulation of gene expression in Caenorhabditis elegans. BMC Genomics 17: 559.
[https://doi.org/10.1186/s12864-016-2837-5]

-
Ceylan, Y., Y. C. Altunoglu and E. Horuz. 2023. HSF and Hsp gene families in sunflower: a comprehensive genome-wide determination survey and expression patterns under abiotic stress conditions. Protoplasma 260(6): 1473-1491.
[https://doi.org/10.1007/s00709-023-01862-6]

-
De Maio, A. 1999. Heat shock proteins: facts, thoughts, and dreams. Shock 11(1): 1-12.
[https://doi.org/10.1097/00024382-199901000-00001]

-
Denlinger, D. L., G. D. Yocum and J. P. Rinehart. 2005. Hormonal control of diapause. pp. 615-650. in Comprehensive Insect Molecular Science, eds. by Gilbert, L. I., K. Iatrou and S. Gill.Elsevier, Amsterdam.
[https://doi.org/10.1016/B0-44-451924-6/00043-0]

-
Dong, D., Y. Zhang, V. Smykal, L. Ling and A. S. Raikhel. 2018. HR38, an ortholog of NR4A family nuclear receptor, mediates 20-hydroxyecdysone regulation of carbohydrate metabolism during mosquito reproduction. Insect Biochem. Mol. Biol. 96: 19-26.
[https://doi.org/10.1016/j.ibmb.2018.02.003]

- FAO. 2017. Food and Agriculture Organization of the United-FAOSTAT-fao.org, .
-
Foottit, R. G. and P. H. Adler. 2017. Insect biodiversity (Science and Society), insect biodiversity informatics. Hoboken, Wiley. pp. 593-602.
[https://doi.org/10.1002/9781118945568.ch18]

-
Gallai, N., J. M. Salles, J. Settele and B. E. Vaissière. 2009. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 68(3): 810-821.
[https://doi.org/10.1016/j.ecolecon.2008.06.014]

-
Garrad, R., D. T. Booth and M. J. Furlong. 2016. The effect of rearing temperature on development, body size, energetics and fecundity of the diamondback moth. Bull. Entomol. Res. 106(2): 175-181.
[https://doi.org/10.1017/S000748531500098X]

- Hilderbrand, M. and G. E. Goslow Jr. 2001. Analysis of vertebrate structure. New York, Wiley. p. 429.
-
Hou, C., H. Rikvin, Y. Slabezki and N. Chehanovsky. 2014. Dynamics of the presence of Israeli acute paralysis virus in honey bee colonies with colony collapse disorder. Viruses 6(5): 2012-2027.
[https://doi.org/10.3390/v6052012]

-
Hu, C., J. Yang, Z. Qi, H. Wu, B. Wang, F. Zou, H. Mei, J. Liu, W. Wang and Q. Liu. 2022. Heat shock proteins: Biological functions, pathological roles, and therapeutic opportunities. MedComm. 3(3): e161.
[https://doi.org/10.1002/mco2.161]

-
Kostál, V. 2006. Eco-physiological phases of insect diapause. J. Insect Physiol. 52(2): 113-127.
[https://doi.org/10.1016/j.jinsphys.2005.09.008]

-
Lautenbach, S., R. Seppelt, J. Liebscher and C. F. Dormann. 2012. Spatial and temporal trends of global pollination benefit. PLoS One 7(4): e35954.
[https://doi.org/10.1371/journal.pone.0035954]

-
Lee Jr., R. E., C. P. Chen and D. L. Denlinger. 1987. A rapid cold-hardening process in insects. Science 238(4832): 1415-1417.
[https://doi.org/10.1126/science.238.4832.1415]

-
Lee, S.-J., S.-H. Kim, J. Lee, J.-H. Kang, S.-M. Lee, H. J. Park, J. Nam and C. Jung. 2022. Impact of ambient temperature variability on the overwintering failure of honeybees in South Korea. J. Apic. 37(3): 331-347.
[https://doi.org/10.17519/apiculture.2022.09.37.3.331]

-
Li, H., X. Zhao, H. Qiao, X. He, J. Tan and D. Hao. 2020. Comparative transcriptome analysis of the heat stress response in Monochamus alternatus Hope (Coleoptera: Cerambycidae). Front. Physiol. 10: 1568.
[https://doi.org/10.3389/fphys.2019.01568]

-
Livak, K. J. and T. D. Schmittgen. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25(4): 402-408.
[https://doi.org/10.1006/meth.2001.1262]

-
Mahanhud, B. and R. Tiwari. 2024. Impact of climate change on honeybees and crop production. in Adapting to climate change in agriculture-theories and practices, eds. by Sheraz, M. S., R. Singh and B. Dhekale. Springer. Cham.
[https://doi.org/10.1007/978-3-031-28142-6_8]

-
Matz, J. M., M. J. Blake, H. M. Tatelman, K. P. Lavoi and N. J. Holbrook. 1995. Characterization and regulation of cold-induced heat shock protein expression in mouse brown adipose tissue. Am. J. Physiol. 269(1 Pt 2): R38-R47.
[https://doi.org/10.1152/ajpregu.1995.269.1.R38]

-
Meiselman, M., S. S. Lee, R. T. Tran, H. Dai, Y. Ding, C. River-Perez, T. P. Wijesekera, B. Dauwalder, F. G. Noriega and A. E. Adams. 2017. Endocrine network essential for reproductive success in Drosophila melanogaster. Proc. Natl. Acad. Sci. USA. 114(19): E3849-E3858.
[https://doi.org/10.1073/pnas.1620760114]

-
Moen, C., J. C. Johnson and J. H. Price. 2022. Ecdysteroid responses to urban heat island conditions during development of the western black widow spider (Latrodectus hesperus). PLoS ONE 17(4): e0267398.
[https://doi.org/10.1371/journal.pone.0267398]

-
Paris, L., M. Roussel, B. Pereiea, F. Delbac and M. Diogon. 2017. Disruption of oxidative balance in the gut of the western honeybee Apis mellifera exposed to the intracellular parasite Nosema ceranae and to the insecticide fipronil. Microb. Biotechnol. 10(6): 1702-1717.
[https://doi.org/10.1111/1751-7915.12772]

-
Richard, D. 1986. Effects of ovarian hormones on energy balance and brown adipose tissue thermogenesis. Am. J. Physiol. 250(2): R245-249.
[https://doi.org/10.1152/ajpregu.1986.250.2.R245]

-
Rosenzweig, R., N. B. Nillegoda, M. P. Mayer and B. Bukau. 2019. The Hsp70 chaperone network. Nat. Rev. Mol. Cell Biol. 20(11): 665-680.
[https://doi.org/10.1038/s41580-019-0133-3]

-
Teets, N. M., A. E. Michael, B. B. Joshua, G. Lopez-Martinez, D. L. Denlinger and R. E. Lee Jr. 2008. Rapid cold-hardening in larvae of the Antarctic midge Belgica antarctica: cellular cold-sensing and a role for calcium. Am. J. Physiol. Regul. Integr. Comp. Physiol. 294(6): R1938-R1946.
[https://doi.org/10.1152/ajpregu.00459.2007]

-
Teets, N. M., S.-X. Yi, R. E. Lee Jr and D. L. Denlinger. 2013. Calcium signaling mediates cold sensing in insect tissue. Proc. Natl. Acad. Sci. USA 110(22): 9154-9159.
[https://doi.org/10.1073/pnas.1306705110]

-
Trenti, F., T. Sandron, G. Guella and V. Lencioni. 2022. Insect cold-tolerance and lipidome: Membrane lipid composition of two chironomid species differently adapted to cold. Cryobiology 106: 84-90.
[https://doi.org/10.1016/j.cryobiol.2022.03.004]

-
Troidl, K., I. Rüding, W. J. Cai, Y. Mücke, L. Grossekettler, I. Piotrowska, H. Apfelbeck, W. Schierling, O. L. Volger, A. J. Horrevoets, K. Grote, T. Schmitz-Rixen, W. Schaper and C. Troidl. 2009. Actin-binding rho activating protein (Abra) is essential for fluid shear stress-induced arteriogenesis. Arterioscler. Thromb. Vasc. Biol. 29(12): 2093-2101.
[https://doi.org/10.1161/ATVBAHA.109.195305]

-
Ulziibayar, D., H. Jang, T. Begna and C. Jung. 2025. Honey bee, Apis mellifera toxicity of five antibiotic formulations used against the fire blight control in apple orchard. J. Apic. 40(2): 105-111.
[https://doi.org/10.17519/apiculture.2025.06.40.2.105]

-
vanEngelsdorp, D., J. D. Evans, C. Saegerman, C. Mullin, E. Haubruge, B. K. Nguyen, M. Frazier, J. Frazier, C. Dianna, Y. Chen, R. Underwood, D. R. Tarpy and J. S. Pettis. 2009. Colony collapse disorder: a descriptive study. PLoS ONE 4(8): e6481.
[https://doi.org/10.1371/journal.pone.0006481]

-
Vatanparast, M. and Y. Park. 2021. Comparative RNA-seq analyses of Solenopsis japonica (Hymenoptera: Formicidae) reveal gene in response to cold stress. Genes 12(10): 1610.
[https://doi.org/10.3390/genes12101610]

-
Vatanparast, M. and Y. Park. 2022. Adaptive temporal variation in cold tolerance of the invasive fall armyworm, Spodoptera frugiperda. Sci. Rep. 14: 2388.
[https://doi.org/10.1038/s41598-022-08174-4]

-
Vatanparast, M., R. T. Puckett, D.-S. Choi and Y. Park. 2021. Comparison of gene expression in the red imported fire ant (Solenopsis invicta) under low and high temperature stress. Sci. Rep. 11: 18442.
[https://doi.org/10.1038/s41598-021-95779-w]

-
Watanabe, Y. and M. Tanaka. 2018. Regulation of autophagy by the heat shock factor 1-mediated stress response pathway. in Heat shock proteins and stress. Heat Shock Proteins, eds. by Asea, A. and P. Kaur. vol 15. Springer, Cham.
[https://doi.org/10.1007/978-3-319-90725-3_9]

-
Yi, S-X. and R. E. Lee Jr. 2004. In vivo and in vitro rapid cold-hardening protects cells from cold-shock injury in the flesh fly. J. Comp. Physiol. B 174(8): 611-615.
[https://doi.org/10.1007/s00360-004-0450-4]
