Facile Isolation of N1,N5,N10-tri-p-coumaroylspermidine from Korean Darae(Actinidia arguta) Bee Pollen Extract
Abstract
Actinidia arguta (A. arguta, Korean darae) is widely distributed as a major nectar and pollen source, and its bee pollen is known to be the most widely produced monofloral bee pollen in Korea. Phenolamides identified in bee pollen have been reported to exhibit beneficial biological functions in humans. Among them, N1,N5,N10-tri-p-coumaroylspermidine (TPCS) has been associated with antioxidant, anti-inflammatory, and photoprotective activities. However, the purification of TPCS has traditionally been carried out through laborious and time-consuming chromatographic procedures, resulting in low yields and limited reproducibility. Herein, a simple and efficient approach was developed for the isolation and identification of TPCS from A. arguta bee pollen. The methanolic extract was fractionated by MPLC, followed by Sephadex purification. The purified compound was identified as TPCS based on UPLC, LC-Q-TOF/MS, and NMR analyses. Furthermore, to explore the potential biological relevance of TPCS, molecular docking analysis was performed against polo-like kinase 1 (PLK1), which showed a stable binding interaction. This work provides new insights into the phenolamide composition of A. arguta bee pollen and establishes a practical method for the efficient isolation of TPCS from natural sources, thereby supporting future biochemical and pharmacological studies.
Keywords:
Actinidia arguta, Bee pollen, Isolation, Identification, Phenolamide, N1,N5,N10-tri-p-coumaroylspermidine (TPCS), Polo-like kinase 1, Molecular dockingINTRODUCTION
Bee pollen is one of the most fascinating natural materials due to its unique chemical composition and complex mixture of bioactive compounds. It contains abundant nutrients and diverse secondary metabolites such as phenolic acids, flavonoids, and phenolamides (Campos et al., 2010; Komosinska-Vassev et al., 2015; Denisow and Denisow-Pietrzyk, 2016; Abdelnour et al., 2019; Tutun et al., 2021; Maurya et al., 2025). These chemical components offer great potential as functional ingredients in food, pharmaceutical, and nutraceutical applications (Rodríguez-Pólit et al., 2023; Scarselli et al., 2023; Garofalo et al., 2024; Nems, 2025).
Recently, several research groups have reported that the chemical composition of bee pollen has been extensively investigated through the isolation and structural elucidation of compounds from monofloral sources (Jiangtao et al., 2023; Kostić et al., 2023; Gercek et al., 2024; Bi et al., 2025; Talemi et al., 2025). These studies have shown that bee pollen is rich in polyamines conjugated with hydroxycinnamic acids, known as hydroxycinnamic acid amides (HCAAs), which exhibit structural diversity and biological significance (Ulusoy and Kolayli, 2013; Mohdaly et al., 2015; Kieliszek et al., 2017; Abdelnour et al., 2019; El Ghouizi et al., 2020).
Among these compounds, HCAAs such as N1,N5,N10-tri-p-coumaroylspermidine (TPCS) have been particularly emphasized due to their wide distribution in plant reproductive organs and their important physiological roles (Larbi, 2020; Qiao, 2024; Watanabe et al., 2024; Cetinbas-Genc et al., 2025). TPCS is predominantly detected in the pollen grains and anthers of flowering plants, where it contributes to pollen wall stabilization, UV protection, and defense against oxidative stress (Denisow and Denisow-Pietrzyk, 2016; Rodríguez-Pólit et al., 2023). In addition, TPCS and related HCAAs are known to exhibit antioxidant, anti-inflammatory, and photoprotective activities (Ulusoy and Kolayli, 2013; Komosinska-Vassev et al., 2015; Mohdaly et al., 2015; Kieliszek et al., 2017; Nascimento and Luz, 2018; Gercek et al., 2024).
Despite the growing interest in the bioactivities of TPCS, its isolation and purification have often been conducted through laborious and time-consuming procedures involving multiple chromatographic steps (Ulusoy and Kolayli, 2013; Mohdaly et al., 2015; Zhuge et al., 2024; Nouri et al., 2025). These conventional methods have typically yielded low quantities of the purified compound, restricting its broader application in biochemical and pharmacological studies (Qiao, 2024; Watanabe et al., 2024; Bi et al., 2025). In previous studies, the purification of TPCS commonly relied on multi-step column chromatography, including repeated silica gel and Sephadex fractionations, which were complex, time-consuming, and resulted in low yield and poor reproducibility. In contrast, the present study established a simplified and efficient purification strategy using medium-pressure liquid chromatography (MPLC) with a dichloromethane-methanol gradient system, enabling the rapid isolation of TPCS with improved yield and reproducibility (Kostić et al., 2023; Scarselli et al., 2023; Garofalo et al., 2024; Bi et al., 2025; Talemi et al., 2025).
In this study, a straightforward isolation route was developed for the identification of TPCS from A. arguta bee pollen extract. The method involves the preparation of a methanol extract from A. arguta bee pollen, followed by fractionation using MPLC, and purification of the dichloromethane (80%) : methanol (20%) fraction to yield TPCS. The compound was identified by ultra-performance liquid chromatography (UPLC), LC-Q-TOF/MS, and NMR analyses. Furthermore, to elucidate the potential biological relevance of TPCS, molecular docking analysis was performed against polo-like kinase 1 (PLK1), a key regulatory enzyme involved in cell-cycle progression and cellular proliferation (Celhar et al., 2016; Zhuge et al., 2024).
MATERIALS AND METHODS
1. Bee pollen samples
Bee pollen of A. arguta was purchased offline in May 2024 in the Republic of Korea. The authenticity of the sample was verified based on its floral morphology and the regional flowering period of A. arguta. After purchase, the pollen was freeze-dried at -4℃ for 24 h to remove water. The dried pollen was stored in a deep freezer at -80℃
2. Extraction procedure
A. arguta bee pollen (5 g) was dispersed in MeOH 50 mL under continuous sonication and the mixture was stirred for 24 h. The obtained mixture was filtered, and washed with MeOH several times. The combined filtrates were concentrated under reduced pressure at 40℃ using a rotary evaporator, and approximately 2 g of crude extract was obtained.
3. MPLC fractionation
MPLC separations were carried out using a Biotage Selekt system (Biotage, Uppsala, Sweden) equipped with a Biotage Silica column. A total of 2 g of the bee pollen extract was used for the fractionation. DCM and MeOH were used as the mobile phase. The separation was performed under isocratic conditions at a constant flow rate of 120 mL/min. Fractions were collected by gradually decreasing the DCM concentration in steps of 100%, 90%, 80%, 70%, 60%, 50%, 40%, 30%, 20%, and 10%. A total of ten fractions, each of 500 mL, were collected. All fractions were concentrated under reduced pressure and subsequently analyzed using UPLC to determine their chemical composition.
4. Isolation and identification
The fraction that showed a distinct UV absorption peak in the MPLC profile (fraction 3, eluted with 80% DCM and 20% MeOH, 0.4 g) was further purified using an open-column chromatography system packed with Sephadex LH-20. The separation was performed under isocratic conditions with a solvent mixture consisting of 90 mL of methanol (MeOH) and 10 mL of distilled water (9 : 1, v/v). The eluates were monitored by UV absorbance at 254 nm and by reversed-phase thin-layer chromatography (TLC) to track the elution of compounds. The purified compound was subsequently identified based on UPLC, LC-Q-TOF/MS, and nuclear magnetic resonance spectroscopy (1H and 13C NMR) analyses.
5. Molecular docking analysis
Molecular docking analysis was performed to predict the binding affinity and interaction mode between TPCS and PLK1. The crystal structure of PLK1 (PDB ID: 3D5U) was obtained from the RCSB Protein Data Bank (https://www.rcsb.org). The protein consists of 603 amino acid residues with a molecular weight of approximately 68.4 kDa. The protein structure was prepared by removing water molecules and adding hydrogen atoms, while the ligand structure of TPCS was optimized using its energy-minimized conformation. Docking calculations were carried out using AutoDock Vina software (Scripps Research Institute, La Jolla, CA, USA), and the most stable conformation was selected based on the lowest binding energy. The protein-ligand complex was visualized using PyMOL Molecular Graphics System (Schrödinger, LLC, New York, NY, USA) to identify key interacting residues and hydrogen-bonding patterns.
RESULTS AND DISCUSSION
The methanolic extraction of A. arguta bee pollen (5 g) was performed, and 2.0 g of crude extract was obtained (yield=40% w/w). The crude extract was fractionated by MPLC using DCM and MeOH as eluents. Elution was carried out by gradually decreasing the concentration of DCM from 100% to 10% (v/v) in 10% increments, resulting in a total of ten fractions. The individual yields of fractions 1-10 were calculated to be 0.95%, 1.90%, 24.45%, 32.34%, 11.59%, 5.87%, 7.39%, 2.09%, 2.24%, and 0.42%, respectively, corresponding to a total recovery yield of approximately 89.2%. Fig. 1 shows the typical UPLC chromatograms obtained from (a) Fr 1. DCM 100%, (b) Fr 2. DCM 90%, (c) Fr 4. DCM 70%, (d) Fr 5. DCM 60%, (e) Fr 6. DCM 50%, (f) Fr 7. DCM 40%, (g) Fr 8. DCM 30%, (h) Fr 9. DCM 20%, and (i) Fr 10. DCM 10%, respectively.
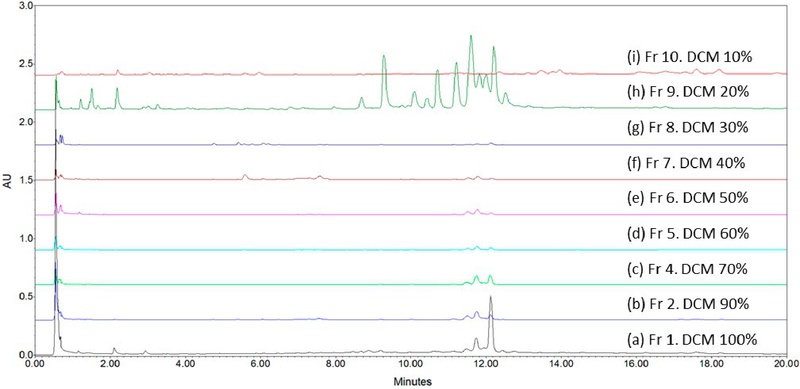
UPLC chromatograms of (a) Fr 1. DCM 100%, (b) Fr 2. DCM 90%, (c) Fr 4. DCM 70%, (d) Fr 5. DCM 60%, (e) Fr 6. DCM 50%, (f) Fr 7. DCM 40%, (g) Fr 8. DCM 30%, (h) Fr 9. DCM 20%, and (i) Fr 10. DCM 10%.
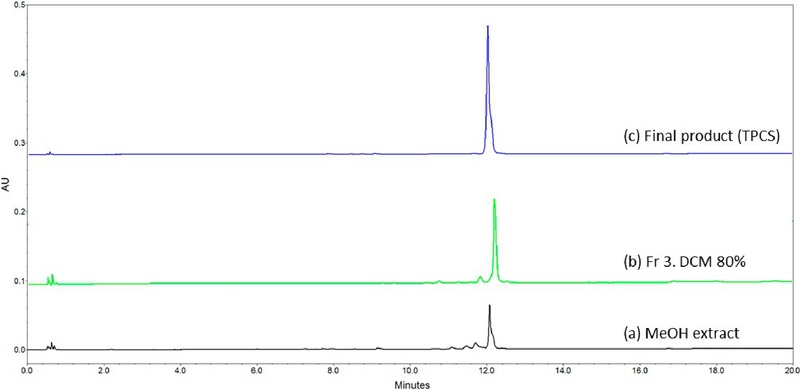
Among these, Fraction 3 (DCM 80%+MeOH 20%) exhibited a distinct UV absorption band at 254 nm and was subsequently purified using a Sephadex LH-20 column. Fig. 2 shows the typical UPLC chromatograms obtained from (a) MeOH extract, (b) Fr 3. DCM 80%, and (c) Final product. After further purification of Fraction 3, 44 mg of purified compound was obtained, corresponding to a yield of approximately 6.8% based on the weight of the original fraction.
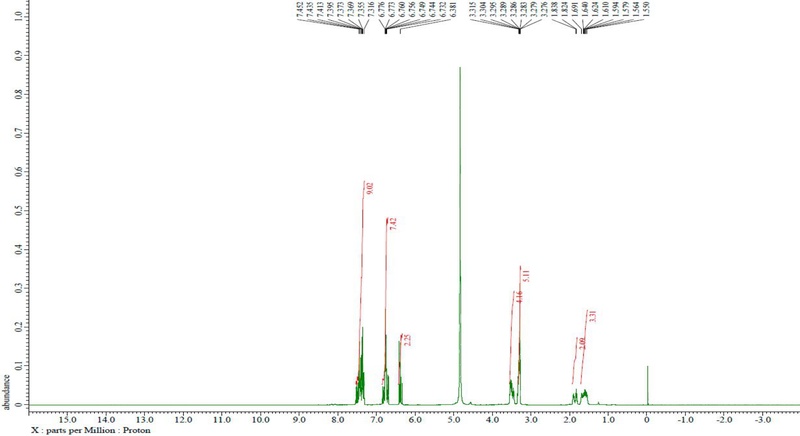
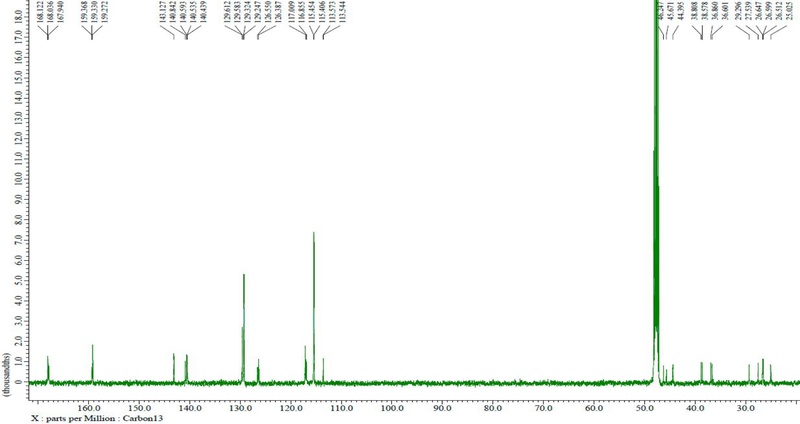
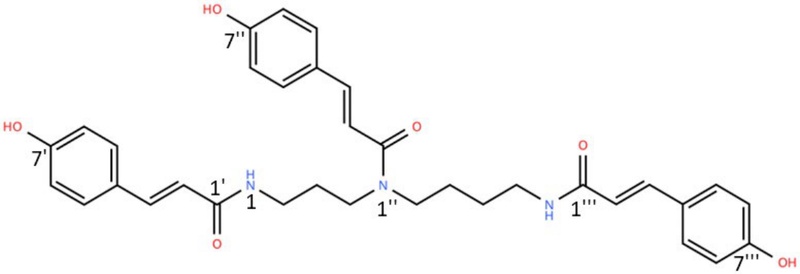
The final compound was identified as TPCS based on LC-Q-TOF/MS and NMR analyses. The major molecular ion peak was observed at m/z 582.2645 ([M-H]-), which was in close agreement with the calculated theoretical value (m/z 582.2604), showing a mass error of 7.04 ppm. Such a small deviation was considered acceptable. The 1H and 13C NMR spectra of the isolated compound are presented in Figs. 3 and 4, respectively.
1H NMR (500 MHz, CD3OD): δH: 1.55 (2H, m, H-8), 1.56 (2H, m, H-7), 1.84 (quin, J=6.8 Hz)/1.92 (quin, J=7.0 Hz) (2H, H-3), 3.28 (4H, m, H-2, 9), 3.31 (4H, m, H-4, 6), 6.38 (d, J=15.0 Hz)/6.40 (d, J=15.0 Hz) (H-8ʹ), 6.73 (d, J=15.5 Hz) (H-8‴), 6.74 (d, J=9.0 Hz)/6.76 (overlapped) (H-3ʺ, 5ʺ), 6.76 (overlapped)/6.77 (overlapped), 6.78 (overlapped) (H-3‴, 5‴), 6.78 (d, J=15.5)/6.78 (d, J=15.5 Hz) (H-8ʺ), 7.31 (d, J=9.0 Hz)/7.35 d, J=9.0 Hz) (H-2ʹ, 6ʹ), 7.31 (d, J=9.0 Hz) (H-2‴, 6‴), 7.31 (overlapped)/7.36 (overlapped)/7.37 (d, J=9.0 Hz) (H-2ʺ, 6ʺ), 7.39 (d. J=15.5 Hz)/7.41 (d, J=15.5 Hz) (H-7‴), 7.43 (d, J=15.5 Hz)/7.45 (d, J=15.5 Hz) (H-7ʺ). 13C-NMR (125 MHz, CD3OD): δH: 25.0/26.5 (C-7), 26.6/26.6 (C-8), 27.5/29.3 (C-3), 36.6/36.9 (C-2), 38.6/38.8 (C-9), 44.4/45.7 (C-4), 45.6/46.2 (C-6), 113.5 (C-8ʺ), 113.6 (C-3ʹ, 3ʺ, 3‴, 5ʹ, 5ʺ, 5‴), 115.4/115.5 (C-8ʹ), 116.8 (C-8‴), 117.0 (C-1ʹ, 1‴), 126.4 (C-1ʺ), 126.6 (C-2ʹ, 6ʹ, 2‴, 6‴), 129.6 (C-2ʺ, 6ʺ), 140.4/140.6/140.8 (C-7ʹ, 7‴), 143.1 (C-7ʺ), 159.3/159.3 (C-4ʹ, 4ʺ, 4‴), 159.4 (C-9ʹ), 167.9/168.0 (C-9ʺ), 168.1 (C-9‴). In comparison with the previously reported data, the LC-TOF-MS and NMR spectral data of the final product were found to be consistent with those of TPCS, as shown in Fig. 5.
Molecular docking analysis was performed to investigate the binding interaction between TPCS and PLK1. TPCS was found to exhibit a strong binding affinity toward PLK1, with a calculated binding energy of -9.0 kca mol-1 and an estimated inhibition constant (Ki) of 252.9 nM. The binding site was predicted to be located within the ATP-binding pocket of PLK1, involving key residues such as Lys82, Leu59, Glu131, Cys133, Phe183, and Asp194. The phenolic hydroxyl groups of TPCS were found to form hydrogen bonds with the backbone atoms of Lys82 and Glu131, whereas the aromatic coumaroyl rings were stabilized through π-π stacking interactions with Phe183. These interactions indicated that TPCS was stably positioned within the hydrophobic cleft of PLK1, suggesting a specific and stable binding mode.
In the Fig. 6, the green ribbon represents the three-dimensional structure of PLK1, while TPCS is displayed as a stick model bound in the active pocket. Yellow dashed lines indicate hydrogen-bond interactions, blue atoms represent nitrogen (N), and red atoms represent oxygen (O) of hydroxyl groups (-OH) involved in hydrogen bonding. Based on these results, it was confirmed that TPCS was bound within the active site of PLK1 through multiple hydrogen-bond and hydrophobic interactions, suggesting its potential role as a functional compound capable of inhibiting PLK1 activity.
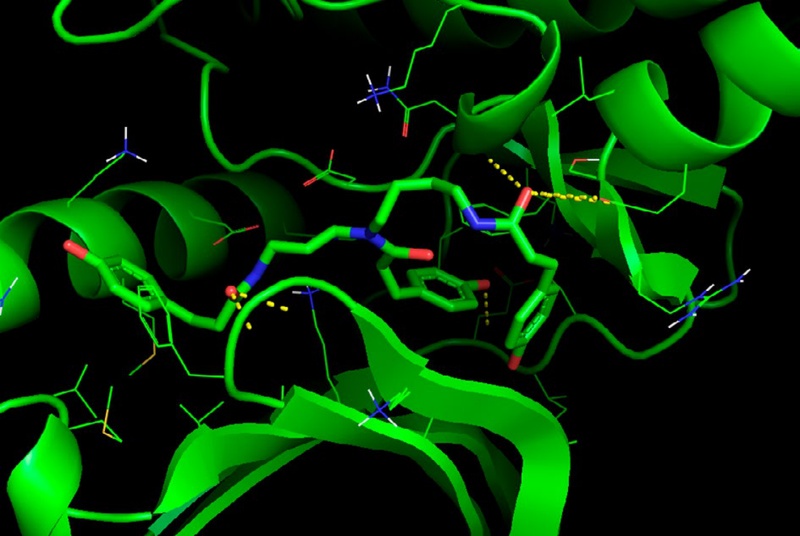
Predicted binding mode of TPCS within PLK1 visualized using PyMOL. The green ribbon represents the PLK1 protein structure, the blue atoms indicate nitrogen (N), the red atoms indicate oxygen (O) of hydroxyl groups (-OH), and the yellow dashed lines indicate hydrogen bonds between TPCS and PLK1 residues.
CONCLUSION
In summary, a facile isolation method for TPCS was successfully established from Actinidia arguta bee pollen extract. The isolated compound was structurally identified by LC-MS and NMR analyses. Through molecular docking analysis, TPCS was found to be stably bound to PLK1, indicating that this phenolamide compound may possess potential biological relevance. The simplified isolation process and molecular characterization presented in this study are expected to provide a practical approach for the further exploration and application of bioactive compounds from bee pollen.
Acknowledgments
This work was carried out with the support of “Cooperative Research Program for Agriculture Science and Technology Development (Project No. RS-2024-00332514)” Rural Development Administration, Republic of Korea.
References
-
Abdelnour, S. A., M. E. Abd El-Hack, M. Alagawany, M. R. Farag and S. S. Elnesr. 2019. Beneficial impacts of bee pollen in animal production, reproduction and health. J. Anim. Physiol. Anim. Nutr. 103: 477-484.
[https://doi.org/10.1111/jpn.13049]

-
Bi, Y., S. Luo, J. Ni, S. Miao, Z. Ning, Z. Zhang, S. Xu, W. Tian, W. Peng and X. Fang. 2025. Effect of ultrasonic treatment on the microstructure, antioxidant activities and metabolites of camellia bee pollen. Ultrason. Sonochem. 118: 107359.
[https://doi.org/10.1016/j.ultsonch.2025.107359]

-
Campos, M. G. R., C. Frigerio, J. Lopes and S. Bogdanov. 2010. What is the future of bee-pollen? J. ApiProd. ApiMed. Sci. 2(4): 131-144.
[https://doi.org/10.3896/IBRA.4.02.4.01]

- Celhar, T., A.-M. Fairhurst and J. E. Allen. 2016. Targeting cyclic GMP-AMP synthase and the STING pathway for the treatment of systemic lupus erythematosus. Expert Rev. Clin. Immunol. 12(10): 1035-1037.
-
Cetinbas-Genc, A., S. Bayraktar and H. Genc. 2025. Pollenkitt from Narcissus tazetta: Evaluation of its UV-protective potential. Chem. Biodivers. 22: e202400621.
[https://doi.org/10.1002/cbdv.202502122]

-
Denisow, B. and M. Denisow-Pietrzyk. 2016. Biological and therapeutic properties of bee pollen: A review. J. Sci. Food Agric. 96: 4303-4309.
[https://doi.org/10.1002/jsfa.7729]

-
El Ghouizi, A., N. El Menyiy, S. I. Falcão, M. Vilas-Boas and B. Lyoussi. 2020. Chemical composition, antioxidant activity, and diuretic effect of Moroccan fresh bee pollen in rats. Vet. World 13(7): 1251-1261.
[https://doi.org/10.14202/vetworld.2020.1251-1261]

-
Garofalo, C., M. Braghini and M. E. Conti. 2024. Bee pollen as a food and feed supplement and a therapeutic remedy. Front. Nutr. 11: 1371672.
[https://doi.org/10.3389/fnut.2024.1371672]

-
Gercek, Y. C., E. Dagsuyu, F. N. Basturk, S. Kırkıncı, N. Yıldırım, G. Kıskanç, B. Özmener, Y. S. Unlu, S. N. Kalkan, K. Boztaş, G. C. Oz, R. Yanardağ, N. E. Bayram, A. Ž. Kostić. 2024. Enzyme inhibitory, physicochemical, and phytochemical properties and botanical sources of honey, bee pollen, bee bread, and propolis obtained from the same apiary. Antioxidants 13: 1483.
[https://doi.org/10.3390/antiox13121483]

-
Jiangtao, Q., Z. Feng, Y. Zhang, X. Xiao, J. Dong, E. Haubruge and H. Zhang. 2023. Phenolamide and flavonoid glycoside profiles of 20 types of monofloral bee pollen. Food Chem. 405: 134800.
[https://doi.org/10.1016/j.foodchem.2022.134800]

- Kieliszek, M., K. Piwowarek, A. M. Kot and S. Blazejak. 2017. The effect of storage conditions on the quality of bee pollen. J. Apic. Sci. 61(1): 123-132.
-
Komosinska-Vassev, K., P. Olczyk, J. Kaźmierczak, Ł. Mencner and K. Olczyk. 2015. Bee pollen: Chemical composition and therapeutic application. Evid. Based Complement. Alternat. Med. 2015: 297425.
[https://doi.org/10.1155/2015/297425]

-
Kostić, A. Z., D. D. Milincic, M. B. Barac, S. P. Stanojevic, U. M. Gasic, Z. L. Tesic and M. B. Pesic. 2023. Monofloral corn poppy bee-collected pollen: A detailed phytochemical profile and assessment of antioxidant properties. Antioxidants 12: 1457.
[https://doi.org/10.3390/antiox12071424]

- Larbi, S. 2020. Differentiation of bee pollen origin through phenolic and volatile profiles. M.Sc. Thesis, Escola Superior Agraria de Braganca, Portugal.
- Maurya, N. K., N. Kushwaha, R. K. Keservani, P. Arya and R. Yadav. 2025. Nutraceutical potential of bee pollen and a natural approach to health and wellness: A comprehensive review. Int. J. Food Sci. Nutr. Res. 9(2): 145-162.
-
Mohdaly, A. A. A., A. A. Mahmoud, M. H. H. Roby, I. Smetanska and M. F. Ramadan. 2015. Phenolic extract from propolis and bee pollen: Composition, antioxidant and antibacterial activities. J. Food Biochem. 39(6): 763-770.
[https://doi.org/10.1111/jfbc.12160]

-
Nascimento, A. M. C.B. and G.E. Luz Jr. 2018. Bee pollen properties: uses and potential pharmacological applications - a review. J. Anal. Pharm. Res. 7(5): 513-515.
[https://doi.org/10.15406/japlr.2018.07.00276]

-
Nems, A. 2025. Bee pollen as a source of phenolic compounds in potato snacks: Chemical composition, antioxidant properties and sensory characteristics. Sci. Rep. 15: 1976.
[https://doi.org/10.1038/s41598-025-09776-4]

- Nouri, A., M. Vazirian, H. R. Monsef Esfahani, M. Sharifzadeh, R. Jahani, M. M. Ardakani, M. Khanavi and M. R. Shams Ardekani. 2025. Hypnotic activity of Carthamus tinctorius petal extracts and its purified components. Res. J. Pharmacogn. 12(1): 1-10.
- Qiao, J. 2024. Phytochemical identification and anti-prostatitis effect of bee pollen. Ph.D. Thesis, University of Liège - Gembloux Agro-Bio Tech, Liège, Belgium.
-
Rodríguez-Pólit, C., R. Gonzalez-Pastor, J. Heredia-Moya, S. E. Carrera-Pacheco, F. Castillo-Solis, R. Vallejo-Imbaquingo, C. Barba-Ostria and L. P. Guamán. 2023. Chemical properties and biological activity of bee pollen. Molecules 28: 7768.
[https://doi.org/10.3390/molecules28237768]

- Scarselli, R., M. Foddai, C. Donati and A. G. Sabatini. 2023. Antioxidant-rich polyfloral bee pollen exerts antimicrobial activity: A case from Tuscany (Italy). Foods 14(5): 802.
-
Talemi, F. P., A. Pourfarzad and S. Gheibi. 2025. Synergistic effects of bee pollen and propolis extract on protein bar properties: A multivariate chemometric analysis. Discover Food 5: 226.
[https://doi.org/10.1007/s44187-025-00541-0]

-
Tutun, H., M. M. Kaya, M. S. Usluer and H. A. Kahraman. 2021. Bee pollen: Its antioxidant activity. Uludag Bee J. 21(1): 119-131.
[https://doi.org/10.31467/uluaricilik.896045]

-
Ulusoy, E. and S. Kolayli. 2013. Phenolic composition and antioxidant properties of Anzer bee pollen. J. Food Biochem. 38(1): 73-82.
[https://doi.org/10.1111/jfbc.12027]

-
Watanabe, C., A. Yanagihara, R. Miyata, T. Mitsui, C. Honda, D. Fujinami and S. Kumazawa. 2024. Catechol-O-methyltransferase and monoamine oxidase B inhibitory activities of Australian bee pollen. Biosci. Biotechnol. Biochem. 88: 665-670.
[https://doi.org/10.1093/bbb/zbae041]

-
Zhuge, H., Z. Ge, J. Wang, J. Yao, J. He, Y. Wang, Y. Wang and Y. Tang. 2024. The tandem of liquid chromatography and network pharmacology for the chemical profiling of Pule’an tablets and the prediction of mechanism of action in treating prostatitis. Pharmaceuticals 17: 56.
[https://doi.org/10.3390/ph17010056]